
|

|

|
| Information on 1ciy |
| PDB: 1ciy Compound: cryia(a) Classification: TOXIN Entry date in PDB: 1997-01-27 Resolution [Å]: 2.25 R-Factor: 0.163 |
CHAIN: - SWISS-PROT/TREMBL:
P02965
KEYWORD: 3D-structure Plasmid Sporulation Toxin SCOP: b.18.1.3 All beta proteins Galactose-binding domain-like Galactose-binding domain-like delta-Endotoxin, C-terminal domain delta-Endotoxin, C-terminal domain Bacillus thuringiensis, CRYIA (A) SCOP: b.77.2.1 All beta proteins beta-Prism I delta-Endotoxin (insectocide), middle domain delta-Endotoxin (insectocide), middle domain delta-Endotoxin (insectocide), middle domain Bacillus thuringiensis, CRYIA (A) SCOP: f.1.3.1 Membrane and cell surface proteins and peptides Toxins' membrane translocation domains delta-Endotoxin (insectocide), N-terminal domain delta-Endotoxin (insectocide), N-terminal domain delta-Endotoxin (insectocide), N-terminal domain Bacillus thuringiensis, CRYIA (A) GO: receptor binding defense response pathogenesis |
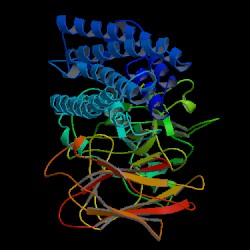
Image Source: PDB |
| Stored Loops of 1ciy |
Loops in ArchDB40 clusters
1ciy_*_480 - AR => SUBCLASS : 3.1.1
1ciy_*_522 - AR => SUBCLASS : 3.2.5
1ciy_*_522 - AR => SUBCLASS : 3.2.6
1ciy_*_522 - AR => SUBCLASS : 3.2.14
1ciy_*_580 - AR => SUBCLASS : 3.3.5
1ciy_*_464 - AR => SUBCLASS : 3.5.4
1ciy_*_470 - AR => SUBCLASS : 6.5.1
1ciy_*_90 - HH => SUBCLASS : 4.3.1
1ciy_*_70 - HH => SUBCLASS : 5.1.2
Loops in ArchDB95 clusters
1ciy_*_480 - AR => SUBCLASS : 1.1.1
1ciy_*_522 - AR => SUBCLASS : 3.1.10
1ciy_*_480 - AR => SUBCLASS : 3.3.3
1ciy_*_464 - AR => SUBCLASS : 3.5.5
1ciy_*_464 - AR => SUBCLASS : 3.6.5
1ciy_*_464 - AR => SUBCLASS : 3.6.7
1ciy_*_470 - AR => SUBCLASS : 5.5.1
1ciy_*_259 - AR => SUBCLASS : 5.46.1
1ciy_*_154 - HH => SUBCLASS : 3.2.8
1ciy_*_90 - HH => SUBCLASS : 4.2.1
1ciy_*_70 - HH => SUBCLASS : 5.1.3
Loops not clustered in ArchDB
1ciy_*_505 - AR
1ciy_*_452 - AR
1ciy_*_446 - AR
1ciy_*_543 - AR
1ciy_*_429 - AR
1ciy_*_400 - AR
1ciy_*_566 - AR
1ciy_*_345 - AR
1ciy_*_338 - AR
1ciy_*_333 - AR
1ciy_*_498 - AR
1ciy_*_266 - EH
1ciy_*_408 - EH
1ciy_*_534 - HA
1ciy_*_298 - HA
1ciy_*_486 - HA
1ciy_*_437 - HA
1ciy_*_380 - HA
1ciy_*_357 - HA
1ciy_*_313 - HA
1ciy_*_423 - HE
1ciy_*_284 - HE
1ciy_*_241 - HE
1ciy_*_35 - HH
1ciy_*_54 - HH
1ciy_*_124 - HH
1ciy_*_186 - HH
1ciy_*_223 - HH
1ciy_*_271 - HH