
|

|

|
| Information on 1cr1 |
| PDB: 1cr1 Compound: DNA primaseHELICASE Classification: TRANSFERASE Entry date in PDB: 1999-11-10 Resolution [Å]: 2.30 R-Factor: 0.235 |
CHAIN: A SWISS-PROT/TREMBL:
P03692
KEYWORD: 3D-structure Alternative initiation ATP-binding DNA replication DNA-directed RNA polymerase Helicase Late protein Metal-binding Primosome Transferase Zinc-finger EC: 2.7.7.- SCOP: c.37.1.11 Alpha and beta proteins (a/b) P-loop containing nucleoside triphosphate hydrolases P-loop containing nucleoside triphosphate hydrolases RecA protein-like (ATPase-domain) Gene 4 protein (g4p, DNA primase), helicase domain Bacteriophage T7 GO: DNA primase activity nucleic acid binding DNA modification DNA replication, synthesis of RNA primer |
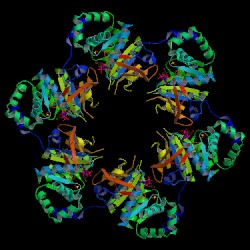
Image Source: PDB |
| Stored Loops of 1cr1 |
Loops in ArchDB40 clusters
1cr1_A_440 - HE => SUBCLASS : 1.1.1
1cr1_A_318 - HE => SUBCLASS : 2.1.1
1cr1_A_490 - HE => SUBCLASS : 2.3.1
1cr1_A_402 - HE => SUBCLASS : 3.1.1
1cr1_A_318 - HE => SUBCLASS : 3.2.3
1cr1_A_528 - HA => SUBCLASS : 4.1.2
1cr1_A_528 - HA => SUBCLASS : 6.1.1
1cr1_A_337 - EH => SUBCLASS : 5.1.1
1cr1_A_337 - EH => SUBCLASS : 5.11.2
1cr1_A_307 - EH => SUBCLASS : 6.1.1
1cr1_A_366 - HH => SUBCLASS : 1.3.1
1cr1_A_366 - HH => SUBCLASS : 1.6.1
Loops in ArchDB95 clusters
1cr1_A_337 - EH => SUBCLASS : 4.2.2
1cr1_A_307 - EH => SUBCLASS : 6.1.1
1cr1_A_337 - EH => SUBCLASS : 6.13.2
1cr1_A_440 - HE => SUBCLASS : 1.1.1
1cr1_A_318 - HE => SUBCLASS : 2.1.1
1cr1_A_490 - HE => SUBCLASS : 2.2.1
1cr1_A_402 - HE => SUBCLASS : 3.1.1
1cr1_A_377 - HE => SUBCLASS : 4.11.2
1cr1_A_528 - HA => SUBCLASS : 4.1.1
Loops not clustered in ArchDB
1cr1_A_391 - EH
1cr1_A_420 - EH
1cr1_A_459 - EH
1cr1_A_498 - HA
1cr1_A_514 - HA
1cr1_A_295 - HE
1cr1_A_272 - HH
1cr1_A_346 - HH
| Homologous structures to 1cr1 classified in ArchDB |
1cr2 A - percentage of sequence identity: 100
1cr4 A - percentage of sequence identity: 100
1e0j A - percentage of sequence identity: 99
1e0j B - percentage of sequence identity: 99
1e0j C - percentage of sequence identity: 99
1e0j D - percentage of sequence identity: 99
1e0j E - percentage of sequence identity: 99
1e0j F - percentage of sequence identity: 99
1e0k A - percentage of sequence identity: 99
1e0k B - percentage of sequence identity: 99
1e0k C - percentage of sequence identity: 99
1e0k D - percentage of sequence identity: 99
1e0k E - percentage of sequence identity: 99
1e0k F - percentage of sequence identity: 99
1q57 A - percentage of sequence identity: 98
1q57 B - percentage of sequence identity: 98
1q57 C - percentage of sequence identity: 98
1q57 D - percentage of sequence identity: 98
1q57 E - percentage of sequence identity: 98
1q57 F - percentage of sequence identity: 98
1q57 G - percentage of sequence identity: 98