
|

|

|
| Information on 1h16 |
| PDB: 1h16 Compound: formate acetyltransferase 1 Classification: TRANSFERASE Entry date in PDB: 2002-11-01 Resolution [Å]: 1.53 R-Factor: 0.145 |
CHAIN: A SWISS-PROT/TREMBL:
P09373
KEYWORD: 3D-structure Acetylation Acyltransferase Complete proteome Glucose metabolism Organic radical Transferase EC: 2.3.1.54 SCOP: c.7.1.1 Alpha and beta proteins (a/b) PFL-like glycyl radical enzymes PFL-like glycyl radical enzymes Pyruvate formate-lyase, PFL Pyruvate formate-lyase, PFL Escherichia coli GO: cytoplasm catalytic activity formate C-acetyltransferase activity carbohydrate metabolism glucose metabolism metabolism |
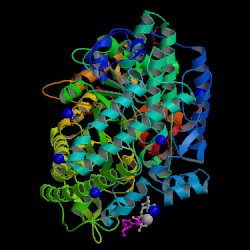
Image Source: PDB |
| Stored Loops of 1h16 |
Loops in ArchDB40 clusters
1h16_A_414 - HA => SUBCLASS : 3.16.1
1h16_A_543 - HA => SUBCLASS : 4.2.2
1h16_A_398 - EH => SUBCLASS : 0.1.41
1h16_A_666 - EH => SUBCLASS : 1.1.1
1h16_A_704 - EH => SUBCLASS : 1.1.5
1h16_A_519 - EH => SUBCLASS : 1.4.1
1h16_A_704 - EH => SUBCLASS : 3.5.5
1h16_A_370 - EH => SUBCLASS : 4.2.2
1h16_A_736 - EH => SUBCLASS : 5.2.1
1h16_A_736 - EH => SUBCLASS : 5.2.2
1h16_A_182 - HH => SUBCLASS : 1.4.1
1h16_A_144 - HH => SUBCLASS : 2.1.1
1h16_A_131 - HH => SUBCLASS : 3.13.2
1h16_A_280 - HH => SUBCLASS : 4.1.1
1h16_A_280 - HH => SUBCLASS : 4.1.2
1h16_A_280 - HH => SUBCLASS : 4.4.1
1h16_A_116 - HH => SUBCLASS : 5.1.3
1h16_A_116 - HH => SUBCLASS : 5.11.1
Loops in ArchDB95 clusters
1h16_A_398 - EH => SUBCLASS : 0.1.51
1h16_A_666 - EH => SUBCLASS : 1.1.2
1h16_A_704 - EH => SUBCLASS : 1.1.3
1h16_A_666 - EH => SUBCLASS : 1.1.17
1h16_A_398 - EH => SUBCLASS : 1.1.18
1h16_A_519 - EH => SUBCLASS : 1.4.1
1h16_A_370 - EH => SUBCLASS : 4.2.1
1h16_A_736 - EH => SUBCLASS : 5.2.1
1h16_A_182 - HH => SUBCLASS : 1.4.1
1h16_A_144 - HH => SUBCLASS : 2.1.1
1h16_A_131 - HH => SUBCLASS : 3.19.1
1h16_A_280 - HH => SUBCLASS : 4.1.1
1h16_A_116 - HH => SUBCLASS : 5.1.2
1h16_A_529 - HE => SUBCLASS : 0.1.4
1h16_A_529 - HE => SUBCLASS : 0.1.7
1h16_A_508 - HE => SUBCLASS : 3.27.1
1h16_A_543 - HA => SUBCLASS : 3.1.2
1h16_A_414 - HA => SUBCLASS : 3.17.1
1h16_A_543 - HA => SUBCLASS : 5.1.1
Loops in ArchDB-EC clusters
1h16_A_116 - HH => SUBCLASS : 5.1.2
Loops not clustered in ArchDB
1h16_A_662 - AR
1h16_A_662 - AR
1h16_A_106 - EH
1h16_A_598 - EH
1h16_A_554 - EH
1h16_A_428 - EH
1h16_A_335 - EH
1h16_A_72 - HA
1h16_A_421 - HA
1h16_A_729 - HA
1h16_A_72 - HA
1h16_A_414 - HA
1h16_A_421 - HA
1h16_A_543 - HA
1h16_A_729 - HA
1h16_A_712 - HE
1h16_A_678 - HE
1h16_A_644 - HE
1h16_A_571 - HE
1h16_A_402 - HE
1h16_A_381 - HE
1h16_A_351 - HE
1h16_A_321 - HE
1h16_A_48 - HE
1h16_A_25 - HH
1h16_A_671 - HH
1h16_A_608 - HH
1h16_A_154 - HH
1h16_A_189 - HH
1h16_A_5 - HH
1h16_A_213 - HH
1h16_A_251 - HH
1h16_A_298 - HH
1h16_A_438 - HH
1h16_A_472 - HH
1h16_A_204 - HH
| Homologous structures to 1h16 classified in ArchDB |
1cm5 A - percentage of sequence identity: 99
1cm5 B - percentage of sequence identity: 99
1h17 A - percentage of sequence identity: 100
1h18 A - percentage of sequence identity: 100
1h18 B - percentage of sequence identity: 100
1mzo A - percentage of sequence identity: 100
1mzo B - percentage of sequence identity: 100
1qhm A - percentage of sequence identity: 100
1qhm B - percentage of sequence identity: 100
2pfl A - percentage of sequence identity: 100
2pfl B - percentage of sequence identity: 100
3pfl A - percentage of sequence identity: 100
3pfl B - percentage of sequence identity: 100