
|

|

|
| Information on 1huj |
| PDB: 1huj Compound: inorganic pyrophosphatase Classification: HYDROLASE Entry date in PDB: 1998-04-08 Resolution [Å]: 2.10 R-Factor: 0.209 |
CHAIN: A SWISS-PROT/TREMBL:
P00817
KEYWORD: 3D-structure Hydrolase Magnesium Metal-binding EC: 3.6.1.1 SCOP: b.40.5.1 All beta proteins OB-fold Inorganic pyrophosphatase Inorganic pyrophosphatase Inorganic pyrophosphatase Baker's yeast (Saccharomyces cerevisiae) GO: cytoplasm inorganic diphosphatase activity magnesium ion binding pyrophosphatase activity metabolism CHAIN: B SWISS-PROT/TREMBL:
P00817
KEYWORD: 3D-structure Hydrolase Magnesium Metal-binding EC: 3.6.1.1 SCOP: b.40.5.1 All beta proteins OB-fold Inorganic pyrophosphatase Inorganic pyrophosphatase Inorganic pyrophosphatase Baker's yeast (Saccharomyces cerevisiae) GO: cytoplasm inorganic diphosphatase activity magnesium ion binding pyrophosphatase activity metabolism |
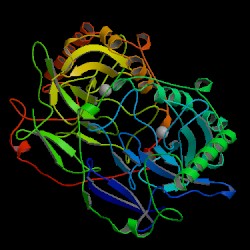
Image Source: PDB |
| Homologous structures to 1huj classified in ArchDB |
1e9g A - percentage of sequence identity: 99