
|

|

|
| Information on 1nhk |
| PDB: 1nhk Compound: Nucleoside diphosphate kinase complexed with 5'-cy Classification: PHOSPHOTRANSFERASE Entry date in PDB: 1995-03-31 Resolution [Å]: 1.90 R-Factor: 0.174 |
CHAIN: L SWISS-PROT/TREMBL:
P15266
KEYWORD: 3D-structure ATP-binding Kinase Transferase EC: 2.7.4.6 SCOP: d.58.6.1 Alpha and beta proteins (a+b) Ferredoxin-like Nucleoside diphosphate kinases Nucleoside diphosphate kinases Nucleoside diphosphate kinases Myxococcus xanthus GO: ATP binding nucleoside-diphosphate kinase activity CTP biosynthesis GTP biosynthesis UTP biosynthesis CHAIN: R SWISS-PROT/TREMBL:
P15266
KEYWORD: 3D-structure ATP-binding Kinase Transferase EC: 2.7.4.6 SCOP: d.58.6.1 Alpha and beta proteins (a+b) Ferredoxin-like Nucleoside diphosphate kinases Nucleoside diphosphate kinases Nucleoside diphosphate kinases Myxococcus xanthus GO: ATP binding nucleoside-diphosphate kinase activity CTP biosynthesis GTP biosynthesis UTP biosynthesis |
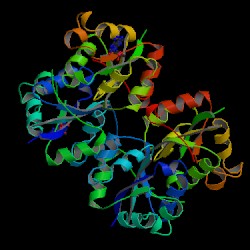
Image Source: PDB |
| Stored Loops of 1nhk |
Loops in ArchDB95 clusters
1nhk_L_33 - EH => SUBCLASS : 3.1.7
1nhk_L_33 - EH => SUBCLASS : 3.1.8
1nhk_L_116 - EH => SUBCLASS : 3.10.2
1nhk_L_44 - HH => SUBCLASS : 1.1.22
1nhk_L_12 - HH => SUBCLASS : 1.3.2
1nhk_L_82 - HH => SUBCLASS : 12.1.1
1nhk_L_60 - HE => SUBCLASS : 0.1.9
1nhk_L_20 - HE => SUBCLASS : 2.5.2
1nhk_L_60 - HE => SUBCLASS : 3.12.1
Loops in ArchDB-EC clusters
1nhk_L_82 - HH => SUBCLASS : 12.1.1
Loops in ArchDB-KI clusters
1nhk_R_3 - EH => SUBCLASS : 1.1.3
1nhk_R_33 - EH => SUBCLASS : 1.1.4
1nhk_R_20 - HE => SUBCLASS : 2.3.2
1nhk_R_60 - HE => SUBCLASS : 2.5.1
1nhk_R_12 - HH => SUBCLASS : 1.1.1
1nhk_R_44 - HH => SUBCLASS : 1.2.1
1nhk_R_82 - HH => SUBCLASS : 12.1.1
Loops not clustered in ArchDB
1nhk_L_3 - EH
1nhk_L_72 - EH
1nhk_L_103 - HE
Loops not clustered in ArchDB-KI
1nhk_R_116 - EH
1nhk_R_72 - EH
1nhk_R_103 - HE
| Homologous structures to 1nhk classified in ArchDB |
1nhk L - percentage of sequence identity: 100
1nhk R - percentage of sequence identity: 100
1nlk L - percentage of sequence identity: 100
1nlk R - percentage of sequence identity: 100
2nck L - percentage of sequence identity: 100
2nck R - percentage of sequence identity: 100