
|

|

|
| Information on 1p8v |
| PDB: 1p8v Compound: platelet glycoprotein ib alpha chain Classification: MEMBRANE PROTEIN/HYDROLASE Entry date in PDB: 2003-07-22 Resolution [Å]: 2.60 R-Factor: 0.207 |
CHAIN: A SWISS-PROT/TREMBL:
P07359
KEYWORD: 3D-structure Bernard Soulier syndrome Blood coagulation Cell adhesion Disease mutation Glycoprotein Hemostasis Leucine-rich repeat Platelet Polymorphism Repeat Signal Sulfation Transmembrane von Willebrand disease 3D-structure Bernard Soulier syndrome Blood coagulation Cell adhesion Disease mutation Glycoprotein Hemostasis Leucine-rich repeat Platelet Polymorphism Repeat Signal Sulfation Transmembrane von Willebrand disease SCOP: c.10.2.7 Alpha and beta proteins (a/b) Leucine-rich repeat, LRR (right-handed beta-alpha superhelix) L domain-like Ngr ectodomain-like von Willebrand factor binding domain of glycoprotein Ib alpha Human (Homo sapiens) GO: CHAIN: B SWISS-PROT/TREMBL:
P00734
KEYWORD: 3D-structure Acute phase Blood coagulation Calcium-binding Disease mutation Gamma-carboxyglutamic acid Glycoprotein Hydrolase Kringle Plasma Polymorphism Repeat Serine protease Signal Vitamin K Zymogen EC: 3.4.21.5 SCOP: b.47.1.2 All beta proteins Trypsin-like serine proteases Trypsin-like serine proteases Eukaryotic proteases Thrombin Human (Homo sapiens) GO: extracellular region calcium ion binding chymotrypsin activity thrombin activity trypsin activity blood coagulation proteolysis and peptidolysis CHAIN: C SWISS-PROT/TREMBL:
P00734
KEYWORD: 3D-structure Acute phase Blood coagulation Calcium-binding Disease mutation Gamma-carboxyglutamic acid Glycoprotein Hydrolase Kringle Plasma Polymorphism Repeat Serine protease Signal Vitamin K Zymogen EC: 3.4.21.5 SCOP: b.47.1.2 All beta proteins Trypsin-like serine proteases Trypsin-like serine proteases Eukaryotic proteases Thrombin Human (Homo sapiens) GO: extracellular region calcium ion binding chymotrypsin activity thrombin activity trypsin activity blood coagulation proteolysis and peptidolysis |
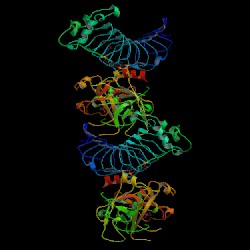
Image Source: PDB |
| Homologous structures to 1p8v classified in ArchDB |
1etr H - percentage of sequence identity: 87
1h8d H - percentage of sequence identity: 97
1jou B - percentage of sequence identity: 99
1ucy H - percentage of sequence identity: 85
1vr1 H - percentage of sequence identity: 96
2hnt F - percentage of sequence identity: 100
null nu - percentage of sequence identity: 0