
|

|

|
| Information on 1pmt |
| PDB: 1pmt Compound: glutathione transferase Classification: TRANSFERASE Entry date in PDB: 1999-04-20 Resolution [Å]: 2.50 R-Factor: 0.202 |
CHAIN: - SWISS-PROT/TREMBL:
P15214
KEYWORD: 3D-structure Transferase EC: 2.5.1.18 SCOP: a.45.1.1 All alpha proteins Glutathione S-transferase (GST), C-terminal domain Glutathione S-transferase (GST), C-terminal domain Glutathione S-transferase (GST), C-terminal domain Class beta GST Proteus mirabilis SCOP: c.47.1.5 Alpha and beta proteins (a/b) Thioredoxin fold Thioredoxin-like Glutathione S-transferase (GST), N-terminal domain Class beta GST Proteus mirabilis GO: |
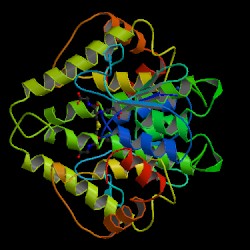
Image Source: PDB |
| Stored Loops of 1pmt |
Loops in ArchDB95 clusters
1pmt_*_176 - HH => SUBCLASS : 1.1.1
1pmt_*_176 - HH => SUBCLASS : 1.1.2
1pmt_*_122 - HH => SUBCLASS : 10.7.1
1pmt_*_66 - HH => SUBCLASS : 12.5.1
1pmt_*_12 - HE => SUBCLASS : 3.5.1
1pmt_*_54 - HA => SUBCLASS : 3.1.1
1pmt_*_26 - HA => SUBCLASS : 4.1.1
Loops in ArchDB-EC clusters
1pmt_*_122 - HH => SUBCLASS : 10.12.1
Loops not clustered in ArchDB
1pmt_*_36 - AR
1pmt_*_36 - AR
1pmt_*_2 - EH
1pmt_*_62 - EH
1pmt_*_26 - HA
1pmt_*_54 - HA
1pmt_*_89 - HH
1pmt_*_106 - HH
1pmt_*_153 - HH
| Homologous structures to 1pmt classified in ArchDB |
2pmt A - percentage of sequence identity: 100
2pmt B - percentage of sequence identity: 100
2pmt C - percentage of sequence identity: 100
2pmt D - percentage of sequence identity: 100