
|

|

|
| Information on 1q1a |
| PDB: 1q1a Compound: hst2 protein Classification: GENE REGULATION Entry date in PDB: 2003-11-18 Resolution [Å]: 1.50 R-Factor: 0.183 |
CHAIN: A SWISS-PROT/TREMBL:
P53686
KEYWORD: Hydrolase Metal-binding NAD Nuclear protein Repressor Transcription regulation Zinc EC: 3.5.1.- SCOP: c.31.1.5 Alpha and beta proteins (a/b) DHS-like NAD/FAD-binding domain DHS-like NAD/FAD-binding domain Sir2 family of transcriptional regulators Hst2 Baker's yeast (Saccharomyces cerevisiae) GO: chromatin silencing complex DNA binding chromatin silencing regulation of transcription, DNA-dependent CHAIN: B SWISS-PROT/TREMBL:
P02309
KEYWORD: 3D-structure Acetylation Chromosomal protein DNA-binding Nuclear protein Nucleosome core GO: |
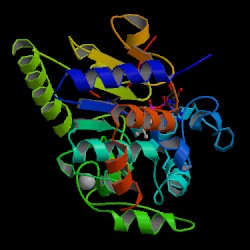
Image Source: PDB |
| Stored Loops of 1q1a |
Loops in ArchDB95 clusters
1q1a_A_264 - EH => SUBCLASS : 1.1.3
1q1a_A_149 - EH => SUBCLASS : 1.1.11
1q1a_A_264 - EH => SUBCLASS : 3.7.2
1q1a_A_264 - EH => SUBCLASS : 3.7.11
1q1a_A_109 - EH => SUBCLASS : 5.77.1
1q1a_A_218 - EH => SUBCLASS : 8.20.1
1q1a_A_69 - HH => SUBCLASS : 1.2.4
1q1a_A_69 - HH => SUBCLASS : 1.2.6
1q1a_A_270 - HH => SUBCLASS : 1.3.3
1q1a_A_95 - HE => SUBCLASS : 2.4.1
1q1a_A_233 - HE => SUBCLASS : 5.1.2
1q1a_A_120 - HE => SUBCLASS : 6.2.8
1q1a_A_136 - HA => SUBCLASS : 4.1.3
1q1a_A_136 - HA => SUBCLASS : 4.1.10
1q1a_A_136 - HA => SUBCLASS : 5.5.2
Loops in ArchDB-EC clusters
1q1a_A_270 - HH => SUBCLASS : 1.3.2
Loops not clustered in ArchDB
1q1a_A_27 - EH
1q1a_A_177 - EH
1q1a_A_243 - EH
1q1a_A_131 - HA
1q1a_A_131 - HA
1q1a_A_136 - HA
1q1a_A_254 - HE
1q1a_A_152 - HE
1q1a_A_9 - HE
1q1a_A_190 - HE
1q1a_A_77 - HH
| Homologous structures to 1q1a classified in ArchDB |
1q17 A - percentage of sequence identity: 100
1q17 B - percentage of sequence identity: 100
1q17 C - percentage of sequence identity: 100
1szc A - percentage of sequence identity: 100
1szd A - percentage of sequence identity: 100