
|

|

|
| Information on 1smd |
| PDB: 1smd Compound: amylase Classification: HYDROLASE (O-GLYCOSYL) Entry date in PDB: 1996-07-11 Resolution [Å]: 1.60 R-Factor: 0.184 |
CHAIN: - SWISS-PROT/TREMBL:
P04745
KEYWORD: 3D-structure Calcium-binding Carbohydrate metabolism Chloride Glycoprotein Glycosidase Hydrolase Multigene family Pyrrolidone carboxylic acid Signal EC: 3.2.1.1 SCOP: b.71.1.1 All beta proteins Glycosyl hydrolase domain Glycosyl hydrolase domain alpha-Amylases, C-terminal beta-sheet domain Animal alpha-amylase Human (Homo sapiens) SCOP: c.1.8.1 Alpha and beta proteins (a/b) TIM beta/alpha-barrel (Trans)glycosidases Amylase, catalytic domain Animal alpha-amylase Human (Homo sapiens) GO: alpha-amylase activity carbohydrate metabolism |
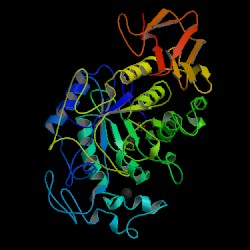
Image Source: PDB |
| Stored Loops of 1smd |
Loops in ArchDB95 clusters
1smd_*_165 - EH => SUBCLASS : 4.1.3
1smd_*_466 - AR => SUBCLASS : 4.13.2
1smd_*_425 - AR => SUBCLASS : 5.1.1
1smd_*_362 - HA => SUBCLASS : 2.1.1
Loops in ArchDB-EC clusters
1smd_*_154 - HE => SUBCLASS : 5.45.1
Loops not clustered in ArchDB
1smd_*_474 - AR
1smd_*_335 - AR
1smd_*_436 - AR
1smd_*_447 - AR
1smd_*_461 - AR
1smd_*_474 - AR
1smd_*_92 - AR
1smd_*_335 - AR
1smd_*_425 - AR
1smd_*_436 - AR
1smd_*_447 - AR
1smd_*_461 - AR
1smd_*_466 - AR
1smd_*_92 - AR
1smd_*_12 - EH
1smd_*_366 - EH
1smd_*_292 - EH
1smd_*_252 - EH
1smd_*_193 - EH
1smd_*_101 - EH
1smd_*_39 - EH
1smd_*_457 - HA
1smd_*_416 - HA
1smd_*_406 - HA
1smd_*_362 - HA
1smd_*_229 - HA
1smd_*_229 - HA
1smd_*_406 - HA
1smd_*_416 - HA
1smd_*_457 - HA
1smd_*_173 - HE
1smd_*_76 - HE
1smd_*_256 - HE
1smd_*_318 - HE
1smd_*_21 - HE
1smd_*_389 - HE
1smd_*_204 - HE
| Homologous structures to 1smd classified in ArchDB |
1b2y A - percentage of sequence identity: 96
1bsi - - percentage of sequence identity: 97
1bvn P - percentage of sequence identity: 85
1c8q A - percentage of sequence identity: 100
1cpu A - percentage of sequence identity: 97
1dhk A - percentage of sequence identity: 86
1hny - - percentage of sequence identity: 97
1jfh - - percentage of sequence identity: 86
1jxj A - percentage of sequence identity: 99
1jxk A - percentage of sequence identity: 98
1kb3 A - percentage of sequence identity: 96
1kbb A - percentage of sequence identity: 96
1kbk A - percentage of sequence identity: 96
1kgu A - percentage of sequence identity: 96
1kgw A - percentage of sequence identity: 96
1kgx A - percentage of sequence identity: 96
1kxq A - percentage of sequence identity: 86
1kxq B - percentage of sequence identity: 86
1kxq C - percentage of sequence identity: 86
1kxq D - percentage of sequence identity: 86
1kxt A - percentage of sequence identity: 86
1kxt C - percentage of sequence identity: 86
1kxt E - percentage of sequence identity: 86
1kxv A - percentage of sequence identity: 86
1kxv B - percentage of sequence identity: 86
1mfu A - percentage of sequence identity: 98
1mfv A - percentage of sequence identity: 100
1nm9 A - percentage of sequence identity: 99
1ose - - percentage of sequence identity: 86
1pif - - percentage of sequence identity: 85
1pig - - percentage of sequence identity: 85
1ppi - - percentage of sequence identity: 86
1q4n X - percentage of sequence identity: 99
1u2y A - percentage of sequence identity: 97
1u30 A - percentage of sequence identity: 97
1u33 A - percentage of sequence identity: 97
1ua3 A - percentage of sequence identity: 85
1vah A - percentage of sequence identity: 85
1wo2 A - percentage of sequence identity: 86
1xcw A - percentage of sequence identity: 97
1xcx A - percentage of sequence identity: 97
1xd0 A - percentage of sequence identity: 97
1xd1 A - percentage of sequence identity: 97
1xgz A - percentage of sequence identity: 96
1xh0 A - percentage of sequence identity: 96
1xh1 A - percentage of sequence identity: 96
1xh2 A - percentage of sequence identity: 96
1xv8 A - percentage of sequence identity: 100
1xv8 B - percentage of sequence identity: 100
1z32 X - percentage of sequence identity: 99
2cpu A - percentage of sequence identity: 96
3cpu A - percentage of sequence identity: 96