
|

|

|
| Information on 1znc |
| PDB: 1znc Compound: carbonic anhydrase iv Classification: LYASE Entry date in PDB: 1997-09-17 Resolution [Å]: 2.80 R-Factor: 0.197 |
CHAIN: A SWISS-PROT/TREMBL:
P22748
KEYWORD: 3D-structure GPI-anchor Lipoprotein Lyase Membrane Signal Zinc EC: 4.2.1.1 SCOP: b.74.1.1 All beta proteins Carbonic anhydrase Carbonic anhydrase Carbonic anhydrase Carbonic anhydrase Human (Homo sapiens), isozyme IV GO: carbonate dehydratase activity zinc ion binding one-carbon compound metabolism CHAIN: B SWISS-PROT/TREMBL:
P22748
KEYWORD: 3D-structure GPI-anchor Lipoprotein Lyase Membrane Signal Zinc EC: 4.2.1.1 SCOP: b.74.1.1 All beta proteins Carbonic anhydrase Carbonic anhydrase Carbonic anhydrase Carbonic anhydrase Human (Homo sapiens), isozyme IV GO: carbonate dehydratase activity zinc ion binding one-carbon compound metabolism |
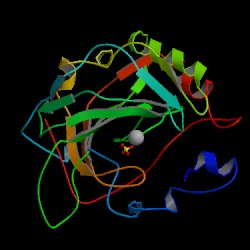
Image Source: PDB |
| Stored Loops of 1znc |
Loops in ArchDB-EC clusters
1znc_A_158 - HE => SUBCLASS : 4.17.2
Loops not clustered in ArchDB
1znc_A_207 - AR
1znc_A_173 - AR
1znc_A_108 - AR
1znc_A_88 - AR
1znc_A_66 - AR
1znc_A_47 - AR
1znc_A_39 - AR
1znc_A_32 - AR
1znc_A_216 - EH
1znc_A_141 - EH
1znc_A_78 - HA
1znc_A_116 - HA
1znc_A_55 - HA
1znc_A_191 - HA
1znc_A_220 - HE
1znc_A_8 - HE
| Homologous structures to 1znc classified in ArchDB |
1znc A - percentage of sequence identity: 100
1znc B - percentage of sequence identity: 100