
|

|

|
| Information on 5csm |
| PDB: 5csm Compound: chorismate mutase Classification: COMPLEX (ISOMERASE/PEPTIDE) Entry date in PDB: 1998-01-14 Resolution [Å]: 2.00 R-Factor: 0.186 |
CHAIN: A SWISS-PROT/TREMBL:
P32178
KEYWORD: 3D-structure Allosteric enzyme Aromatic amino acid biosynthesis Isomerase EC: 5.4.99.5 SCOP: a.130.1.2 All alpha proteins Chorismate mutase II Chorismate mutase II Allosteric chorismate mutase Allosteric chorismate mutase Baker's yeast (Saccharomyces cerevisiae) GO: chorismate mutase activity aromatic amino acid family biosynthesis |
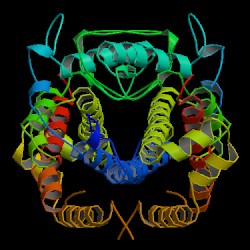
Image Source: PDB |
| Stored Loops of 5csm |
Loops in ArchDB40 clusters
5csm_A_6 - HH => SUBCLASS : 1.1.5
5csm_A_6 - HH => SUBCLASS : 1.1.11
5csm_A_6 - HH => SUBCLASS : 1.1.15
5csm_A_161 - HH => SUBCLASS : 1.2.6
5csm_A_173 - HH => SUBCLASS : 2.2.2
5csm_A_185 - HH => SUBCLASS : 2.3.7
5csm_A_185 - HH => SUBCLASS : 2.3.14
Loops in ArchDB95 clusters
5csm_A_6 - HH => SUBCLASS : 1.1.3
5csm_A_161 - HH => SUBCLASS : 1.2.3
5csm_A_173 - HH => SUBCLASS : 2.2.2
5csm_A_185 - HH => SUBCLASS : 2.3.13
5csm_A_185 - HH => SUBCLASS : 2.4.3
Loops in ArchDB-EC clusters
5csm_A_185 - HH => SUBCLASS : 2.7.1
Loops not clustered in ArchDB
5csm_A_12 - HH
5csm_A_59 - HH
5csm_A_114 - HH
5csm_A_126 - HH
5csm_A_140 - HH
5csm_A_195 - HH
5csm_A_227 - HH
| Homologous structures to 5csm classified in ArchDB |
1csm A - percentage of sequence identity: 98
1csm B - percentage of sequence identity: 98
2csm A - percentage of sequence identity: 98
3csm A - percentage of sequence identity: 99
3csm B - percentage of sequence identity: 99
4csm A - percentage of sequence identity: 98
4csm B - percentage of sequence identity: 98