
|

|

|
| Information on 1akz |
| PDB: 1akz Compound: uracil-DNA glycosylase Classification: GLYCOSIDASE Entry date in PDB: 1997-08-20 Resolution [Å]: 1.57 R-Factor: 0.189 |
CHAIN: - SWISS-PROT/TREMBL:
P13051
KEYWORD: 3D-structure Alternative splicing Disease mutation DNA repair Glycosidase Hydrolase Mitochondrion Nuclear protein Transit peptide EC: 3.2.2.- SCOP: c.18.1.1 Alpha and beta proteins (a/b) DNA glycosylase DNA glycosylase Uracil-DNA glycosylase Uracil-DNA glycosylase Human (Homo sapiens) GO: uracil DNA N-glycosylase activity DNA repair |
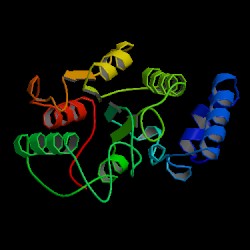
Image Source: PDB |
| Stored Loops of 1akz |
Loops in ArchDB95 clusters
1akz_*_139 - EH => SUBCLASS : 24.1.1
1akz_*_200 - AR => SUBCLASS : 3.4.8
1akz_*_118 - AR => SUBCLASS : 19.2.1
1akz_*_193 - HE => SUBCLASS : 1.2.1
1akz_*_100 - HE => SUBCLASS : 2.6.1
1akz_*_222 - HE => SUBCLASS : 4.56.1
1akz_*_247 - HE => SUBCLASS : 5.13.3
Loops in ArchDB-EC clusters
1akz_*_87 - HH => SUBCLASS : 1.1.1
Loops not clustered in ArchDB
1akz_*_118 - AR
1akz_*_200 - AR
1akz_*_209 - EH
1akz_*_241 - EH
1akz_*_262 - EH
1akz_*_168 - HH
| Homologous structures to 1akz classified in ArchDB |
1emh A - percentage of sequence identity: 100
1emj A - percentage of sequence identity: 100
1okb B - percentage of sequence identity: 76
1q3f A - percentage of sequence identity: 100
1ssp E - percentage of sequence identity: 100
1ugh E - percentage of sequence identity: 100
1yuo A - percentage of sequence identity: 99
2ssp E - percentage of sequence identity: 99
4skn E - percentage of sequence identity: 99