
|

|

|
| Information on 1apm |
| PDB: 1apm Compound: c-AMP-dependent protein kinase (cAPK) (catalytic s Classification: TRANSFERASE(PHOSPHOTRANSFERASE) Entry date in PDB: 1993-04-15 Resolution [Å]: 2.00 R-Factor: 0.186 |
CHAIN: E SWISS-PROT/TREMBL:
P05132
KEYWORD: 3D-structure Alternative splicing ATP-binding cAMP Lipoprotein Multigene family Myristate Nuclear protein Phosphorylation Serine/threonine-protein kinase Transferase EC: 2.7.1.37 SCOP: d.144.1.7 Alpha and beta proteins (a+b) Protein kinase-like (PK-like) Protein kinase-like (PK-like) Protein kinases, catalytic subunit cAMP-dependent PK, catalytic subunit Mouse (Mus musculus) GO: ATP binding protein kinase activity protein serine/threonine kinase activity protein amino acid phosphorylation CHAIN: I SWISS-PROT/TREMBL:
P27776
KEYWORD: 3D-structure Protein kinase inhibitor GO: cAMP-dependent protein kinase inhibitor activity negative regulation of protein kinase activity |
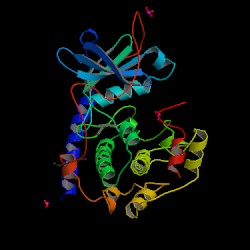
Image Source: PDB |
| Stored Loops of 1apm |
Loops in ArchDB-EC clusters
1apm_E_243 - HH => SUBCLASS : 10.6.1
Loops in ArchDB-KI clusters
1apm_E_180 - AR => SUBCLASS : 5.1.1
1apm_E_162 - AR => SUBCLASS : 8.1.1
1apm_E_68 - EH => SUBCLASS : 0.1.4
1apm_E_115 - EH => SUBCLASS : 6.3.1
1apm_E_43 - HA => SUBCLASS : 2.2.1
1apm_E_106 - HA => SUBCLASS : 2.2.1
1apm_E_172 - HA => SUBCLASS : 4.1.1
1apm_E_55 - HA => SUBCLASS : 4.1.1
1apm_E_140 - HE => SUBCLASS : 1.1.3
1apm_E_85 - HE => SUBCLASS : 8.1.1
1apm_E_11 - HE => SUBCLASS : 11.2.1
1apm_E_76 - HH => SUBCLASS : 1.1.2
1apm_E_128 - HH => SUBCLASS : 4.4.1
1apm_E_289 - HH => SUBCLASS : 8.2.1
1apm_E_218 - HH => SUBCLASS : 9.3.1
1apm_E_243 - HH => SUBCLASS : 11.1.1
1apm_E_263 - HH => SUBCLASS : 14.2.1
Loops not clustered in ArchDB
1apm_E_180 - AR
1apm_E_162 - AR
1apm_E_68 - EH
1apm_E_115 - EH
1apm_E_189 - EH
1apm_E_172 - HA
1apm_E_106 - HA
1apm_E_55 - HA
1apm_E_43 - HA
1apm_E_11 - HE
1apm_E_85 - HE
1apm_E_140 - HE
1apm_E_289 - HH
1apm_E_263 - HH
1apm_E_207 - HH
1apm_E_128 - HH
1apm_E_76 - HH
1apm_E_218 - HH
Loops not clustered in ArchDB-KI
1apm_E_189 - EH
1apm_E_207 - HH
| Homologous structures to 1apm classified in ArchDB |
1atp E - percentage of sequence identity: 99
1bkx A - percentage of sequence identity: 99
1bx6 - - percentage of sequence identity: 99
1cdk A - percentage of sequence identity: 96
1cdk B - percentage of sequence identity: 96
1cmk E - percentage of sequence identity: 96
1ctp E - percentage of sequence identity: 97
1fmo E - percentage of sequence identity: 99
1j3h A - percentage of sequence identity: 99
1j3h B - percentage of sequence identity: 99
1jbp E - percentage of sequence identity: 99
1jlu E - percentage of sequence identity: 99
1l3r E - percentage of sequence identity: 99
1q24 A - percentage of sequence identity: 96
1q62 A - percentage of sequence identity: 96
1q8t A - percentage of sequence identity: 97
1q8u A - percentage of sequence identity: 97
1q8w A - percentage of sequence identity: 97
1re8 A - percentage of sequence identity: 99
1rej A - percentage of sequence identity: 99
1rek A - percentage of sequence identity: 99
1smh A - percentage of sequence identity: 96
1stc E - percentage of sequence identity: 97
1sve A - percentage of sequence identity: 97
1svg A - percentage of sequence identity: 97
1svh A - percentage of sequence identity: 97
1syk A - percentage of sequence identity: 99
1syk B - percentage of sequence identity: 99
1szm A - percentage of sequence identity: 96
1szm B - percentage of sequence identity: 96
1u7e A - percentage of sequence identity: 99
1veb A - percentage of sequence identity: 97
1xh4 A - percentage of sequence identity: 97
1xh5 A - percentage of sequence identity: 97
1xh6 A - percentage of sequence identity: 97
1xh7 A - percentage of sequence identity: 97
1xh8 A - percentage of sequence identity: 97
1xh9 A - percentage of sequence identity: 96
1xha A - percentage of sequence identity: 96
1ydr E - percentage of sequence identity: 97
1yds E - percentage of sequence identity: 97
1ydt E - percentage of sequence identity: 97
2c1a A - percentage of sequence identity: 97
2c1b A - percentage of sequence identity: 97
2cpk E - percentage of sequence identity: 99
2erz E - percentage of sequence identity: 99
null nu - percentage of sequence identity: 0