
|

|

|
| Information on 1c0d |
| PDB: 1c0d Compound: endocellulase e1 from a. cellulolyticus Classification: HYDROLASE Entry date in PDB: 1999-07-23 Resolution [Å]: 2.40 R-Factor: 0.193 |
CHAIN: A SWISS-PROT/TREMBL:
P54583
KEYWORD: 3D-structure Cellulose degradation Glycosidase Hydrolase Signal EC: 3.2.1.4 SCOP: c.1.8.3 Alpha and beta proteins (a/b) TIM beta/alpha-barrel (Trans)glycosidases beta-glycanases Endocellulase E1 Acidothermus cellulolyticus GO: hydrolase activity, hydrolyzing O-glycosyl compounds polysaccharide binding carbohydrate metabolism CHAIN: B SWISS-PROT/TREMBL:
P54583
KEYWORD: 3D-structure Cellulose degradation Glycosidase Hydrolase Signal EC: 3.2.1.4 SCOP: c.1.8.3 Alpha and beta proteins (a/b) TIM beta/alpha-barrel (Trans)glycosidases beta-glycanases Endocellulase E1 Acidothermus cellulolyticus GO: hydrolase activity, hydrolyzing O-glycosyl compounds polysaccharide binding carbohydrate metabolism |
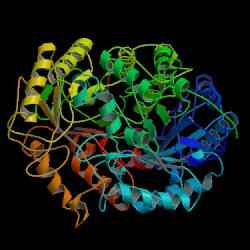
Image Source: PDB |
| Stored Loops of 1c0d |
Loops in ArchDB-EC clusters
1c0d_A_268 - HE => SUBCLASS : 4.2.6
Loops not clustered in ArchDB
1c0d_A_209 - AR
1c0d_A_197 - AR
1c0d_A_20 - AR
1c0d_A_24 - EH
1c0d_A_315 - EH
1c0d_A_278 - EH
1c0d_A_233 - EH
1c0d_A_60 - EH
1c0d_A_110 - EH
1c0d_A_154 - EH
1c0d_A_205 - HA
1c0d_A_12 - HA
1c0d_A_7 - HA
1c0d_A_178 - HE
1c0d_A_306 - HE
1c0d_A_134 - HE
1c0d_A_93 - HE
1c0d_A_46 - HE
1c0d_A_259 - HH
1c0d_A_290 - HH