
|

|

|
| Information on 1cvl |
| PDB: 1cvl Compound: triacylglycerol hydrolase Classification: HYDROLASE Entry date in PDB: 1997-04-01 Resolution [Å]: 1.60 R-Factor: 0.178 |
CHAIN: - SWISS-PROT/TREMBL:
Q05489
KEYWORD: 3D-structure Calcium Hydrolase Lipid degradation Signal EC: 3.1.1.3 SCOP: c.69.1.18 Alpha and beta proteins (a/b) alpha/beta-Hydrolases alpha/beta-Hydrolases Bacterial lipase Lipase Chromobacterium viscosum GO: catalytic activity |
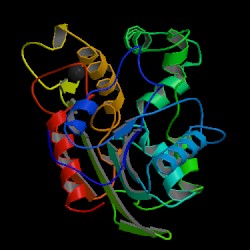
Image Source: PDB |
| Stored Loops of 1cvl |
Loops in ArchDB40 clusters
1cvl_*_33 - HE => SUBCLASS : 2.1.1
1cvl_*_61 - HE => SUBCLASS : 3.1.2
1cvl_*_61 - HE => SUBCLASS : 3.1.4
1cvl_*_89 - HE => SUBCLASS : 4.7.1
1cvl_*_196 - HA => SUBCLASS : 2.4.7
1cvl_*_22 - HA => SUBCLASS : 3.5.1
1cvl_*_81 - EH => SUBCLASS : 1.2.5
1cvl_*_237 - HH => SUBCLASS : 1.2.14
Loops in ArchDB95 clusters
1cvl_*_81 - EH => SUBCLASS : 1.3.7
1cvl_*_237 - HH => SUBCLASS : 1.2.17
1cvl_*_33 - HE => SUBCLASS : 2.1.1
1cvl_*_61 - HE => SUBCLASS : 3.1.1
1cvl_*_61 - HE => SUBCLASS : 3.3.1
1cvl_*_89 - HE => SUBCLASS : 4.6.1
1cvl_*_196 - HA => SUBCLASS : 2.1.6
1cvl_*_22 - HA => SUBCLASS : 3.4.1
Loops in ArchDB-EC clusters
1cvl_*_237 - HH => SUBCLASS : 1.2.3
Loops not clustered in ArchDB
1cvl_*_11 - AR
1cvl_*_202 - AR
1cvl_*_11 - AR
1cvl_*_202 - AR
1cvl_*_27 - EH
1cvl_*_275 - EH
1cvl_*_226 - EH
1cvl_*_104 - EH
1cvl_*_44 - EH
1cvl_*_22 - HA
1cvl_*_214 - HA
1cvl_*_196 - HA
1cvl_*_214 - HA
1cvl_*_268 - HE
1cvl_*_169 - HE
1cvl_*_243 - HH
1cvl_*_137 - HH
1cvl_*_118 - HH
1cvl_*_156 - HH
| Homologous structures to 1cvl classified in ArchDB |
1hqd A - percentage of sequence identity: 84
1oil A - percentage of sequence identity: 84
1oil B - percentage of sequence identity: 84
1qge D - percentage of sequence identity: 100
1qge E - percentage of sequence identity: 98
1tah A - percentage of sequence identity: 100
1tah B - percentage of sequence identity: 100
1tah C - percentage of sequence identity: 100
1tah D - percentage of sequence identity: 100
1ys1 X - percentage of sequence identity: 84
1ys2 X - percentage of sequence identity: 84
2lip - - percentage of sequence identity: 84
3lip - - percentage of sequence identity: 84
4lip E - percentage of sequence identity: 84
5lip - - percentage of sequence identity: 84