
|

|

|
| Information on 1g5t |
| PDB: 1g5t Compound: cob(i)alamin adenosyltransferase Classification: TRANSFERASE Entry date in PDB: 2000-11-22 Resolution [Å]: 1.80 R-Factor: 0.192 |
CHAIN: A SWISS-PROT/TREMBL:
P31570
KEYWORD: 3D-structure ATP-binding Cobalamin biosynthesis Complete proteome Porphyrin biosynthesis Transferase EC: 2.5.1.17 SCOP: c.37.1.11 Alpha and beta proteins (a/b) P-loop containing nucleoside triphosphate hydrolases P-loop containing nucleoside triphosphate hydrolases RecA protein-like (ATPase-domain) ATP:corrinoid adenosyltransferase CobA Salmonella typhimurium GO: ATP binding cob(I)yrinic acid a,c-diamide adenosyltransferase activity cobalamin biosynthesis |
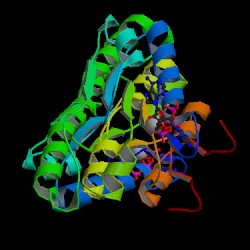
Image Source: PDB |
| Stored Loops of 1g5t |
Loops in ArchDB40 clusters
1g5t_A_72 - HE => SUBCLASS : 1.1.1
1g5t_A_41 - HE => SUBCLASS : 2.1.1
1g5t_A_72 - HE => SUBCLASS : 2.1.9
1g5t_A_165 - HE => SUBCLASS : 2.3.1
1g5t_A_98 - HE => SUBCLASS : 4.5.2
1g5t_A_140 - HE => SUBCLASS : 5.1.1
1g5t_A_123 - EH => SUBCLASS : 1.4.3
1g5t_A_59 - EH => SUBCLASS : 8.9.1
1g5t_A_129 - HH => SUBCLASS : 4.1.3
1g5t_A_129 - HH => SUBCLASS : 4.1.4
Loops in ArchDB95 clusters
1g5t_A_123 - EH => SUBCLASS : 1.4.1
1g5t_A_129 - HH => SUBCLASS : 4.1.1
1g5t_A_72 - HE => SUBCLASS : 1.1.1
1g5t_A_41 - HE => SUBCLASS : 2.1.1
1g5t_A_72 - HE => SUBCLASS : 2.1.21
1g5t_A_165 - HE => SUBCLASS : 2.2.1
1g5t_A_98 - HE => SUBCLASS : 3.4.1
1g5t_A_140 - HE => SUBCLASS : 5.1.2
1g5t_A_98 - HE => SUBCLASS : 5.2.5
1g5t_A_140 - HE => SUBCLASS : 6.41.1
Loops in ArchDB-EC clusters
1g5t_A_129 - HH => SUBCLASS : 4.1.4
Loops not clustered in ArchDB
1g5t_A_31 - EH
1g5t_A_83 - EH
1g5t_A_155 - EH
| Homologous structures to 1g5t classified in ArchDB |
1g5r A - percentage of sequence identity: 100
1g64 A - percentage of sequence identity: 100