
|

|

|
| Information on 1jat |
| PDB: 1jat Compound: ubiquitin-conjugating enzyme e2-17.5 kda Classification: LIGASE Entry date in PDB: 2001-06-20 Resolution [Å]: 1.60 R-Factor: 0.203 |
CHAIN: A SWISS-PROT/TREMBL:
P52490
KEYWORD: 3D-structure Ligase Multigene family Ubl conjugation pathway 3D-structure Ligase Multigene family Ubl conjugation pathway EC: 6.3.2.19 SCOP: d.20.1.1 Alpha and beta proteins (a+b) UBC-like UBC-like Ubiquitin conjugating enzyme, UBC Ubiquitin conjugating enzyme, UBC Baker's yeast (Saccharomyces cerevisiae), ubc13 GO: ubiquitin conjugating enzyme activity protein modification ubiquitin cycle CHAIN: B SWISS-PROT/TREMBL:
P53152
KEYWORD: 3D-structure Ligase Ubl conjugation pathway SCOP: d.20.1.1 Alpha and beta proteins (a+b) UBC-like UBC-like Ubiquitin conjugating enzyme, UBC Ubiquitin conjugating enzyme, UBC Baker's yeast (Saccharomyces cerevisiae), mms2 GO: ubiquitin conjugating enzyme activity protein modification ubiquitin cycle |
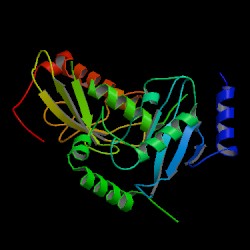
Image Source: PDB |
| Stored Loops of 1jat |
Loops in ArchDB40 clusters
1jat_A_133 - HE => SUBCLASS : 0.1.1
1jat_A_6 - HE => SUBCLASS : 5.13.1
1jat_A_23 - HA => SUBCLASS : 2.2.9
1jat_B_57 - HA => SUBCLASS : 10.4.1
1jat_B_109 - HH => SUBCLASS : 1.1.14
1jat_A_126 - HH => SUBCLASS : 1.2.1
Loops in ArchDB95 clusters
1jat_B_109 - HH => SUBCLASS : 1.1.3
1jat_A_126 - HH => SUBCLASS : 1.2.1
1jat_A_6 - HE => SUBCLASS : 5.17.1
1jat_A_23 - HA => SUBCLASS : 2.2.21
1jat_B_57 - HA => SUBCLASS : 10.2.1
Loops in ArchDB-EC clusters
1jat_A_6 - HE => SUBCLASS : 5.11.1
Loops not clustered in ArchDB
1jat_A_68 - EH
1jat_B_74 - EH
1jat_A_31 - HA
1jat_A_31 - HA
1jat_A_23 - HA
1jat_A_50 - HA
1jat_B_40 - HA
1jat_B_26 - HA
1jat_A_50 - HA
1jat_B_6 - HE
1jat_A_101 - HH
1jat_B_98 - HH
| Homologous structures to 1jat classified in ArchDB |
1jbb A - percentage of sequence identity: 100
1jbb B - percentage of sequence identity: 100
2c2v B - percentage of sequence identity: 69
2c2v E - percentage of sequence identity: 69
2c2v H - percentage of sequence identity: 69
2c2v K - percentage of sequence identity: 69