
|

|

|
| Information on 1qnf |
| PDB: 1qnf Compound: photolyase Classification: DNA REPAIR Entry date in PDB: 1998-01-14 Resolution [Å]: 1.80 R-Factor: 0.197 |
CHAIN: - SWISS-PROT/TREMBL:
P05327
KEYWORD: 3D-structure Chromophore DNA repair DNA-binding FAD Flavoprotein Lyase EC: 4.1.99.3 SCOP: a.99.1.1 All alpha proteins Cryptochrome/photolyase FAD-binding domain Cryptochrome/photolyase FAD-binding domain Cryptochrome/photolyase FAD-binding domain C-terminal domain of DNA photolyase Anacystis nidulans SCOP: c.28.1.1 Alpha and beta proteins (a/b) Cryptochrome/photolyase, N-terminal domain Cryptochrome/photolyase, N-terminal domain Cryptochrome/photolyase, N-terminal domain DNA photolyase Anacystis nidulans GO: DNA photolyase activity DNA repair |
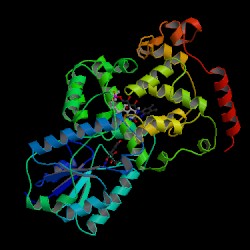
Image Source: PDB |
| Stored Loops of 1qnf |
Loops in ArchDB-EC clusters
1qnf_*_367 - HH => SUBCLASS : 5.2.2
Loops not clustered in ArchDB
1qnf_*_5 - EH
1qnf_*_31 - EH
1qnf_*_73 - EH
1qnf_*_95 - EH
1qnf_*_123 - EH
1qnf_*_104 - HE
1qnf_*_80 - HE
1qnf_*_49 - HE
1qnf_*_19 - HE
1qnf_*_420 - HH
1qnf_*_408 - HH
1qnf_*_383 - HH
1qnf_*_349 - HH
1qnf_*_332 - HH
1qnf_*_318 - HH
1qnf_*_307 - HH
1qnf_*_271 - HH
1qnf_*_254 - HH
1qnf_*_244 - HH
1qnf_*_228 - HH
1qnf_*_210 - HH
1qnf_*_176 - HH
1qnf_*_149 - HH
1qnf_*_39 - HH
1qnf_*_434 - HH
| Homologous structures to 1qnf classified in ArchDB |
1owm A - percentage of sequence identity: 100
1own A - percentage of sequence identity: 100
1owo A - percentage of sequence identity: 100
1owp A - percentage of sequence identity: 100
1tez A - percentage of sequence identity: 100
1tez B - percentage of sequence identity: 100
1tez C - percentage of sequence identity: 100
1tez D - percentage of sequence identity: 100