
|

|

|
| Information on 1s6v |
| PDB: 1s6v Compound: cytochrome c peroxidase, mitochondrial Classification: OXIDOREDUCTASE/ELECTRON TRANSPORT Entry date in PDB: 2004-04-27 Resolution [Å]: 1.88 R-Factor: 0.190 |
CHAIN: A SWISS-PROT/TREMBL:
P00431
KEYWORD: 3D-structure Heme Hydrogen peroxide Iron Mitochondrion Organic radical Oxidoreductase Peroxidase Polymorphism Transit peptide EC: 1.11.1.5 SCOP: a.93.1.1 All alpha proteins Heme-dependent peroxidases Heme-dependent peroxidases CCP-like Cytochrome c peroxidase, CCP Baker's yeast (Saccharomyces cerevisiae) GO: peroxidase activity response to oxidative stress CHAIN: B SWISS-PROT/TREMBL:
P00044
KEYWORD: 3D-structure Electron transport Heme Methylation Mitochondrion Respiratory chain SCOP: a.3.1.1 All alpha proteins Cytochrome c Cytochrome c monodomain cytochrome c Mitochondrial cytochrome c Baker's yeast (Saccharomyces cerevisiae) GO: electron transporter activity heme binding electron transport CHAIN: C SWISS-PROT/TREMBL:
P00431
KEYWORD: 3D-structure Heme Hydrogen peroxide Iron Mitochondrion Organic radical Oxidoreductase Peroxidase Polymorphism Transit peptide EC: 1.11.1.5 SCOP: a.93.1.1 All alpha proteins Heme-dependent peroxidases Heme-dependent peroxidases CCP-like Cytochrome c peroxidase, CCP Baker's yeast (Saccharomyces cerevisiae) GO: peroxidase activity response to oxidative stress CHAIN: D SWISS-PROT/TREMBL:
P00044
KEYWORD: 3D-structure Electron transport Heme Methylation Mitochondrion Respiratory chain SCOP: a.3.1.1 All alpha proteins Cytochrome c Cytochrome c monodomain cytochrome c Mitochondrial cytochrome c Baker's yeast (Saccharomyces cerevisiae) GO: electron transporter activity heme binding electron transport |
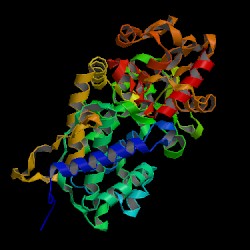
Image Source: PDB |
| Homologous structures to 1s6v classified in ArchDB |
1jdr A - percentage of sequence identity: 97
1ycc - - percentage of sequence identity: 98
1ytc - - percentage of sequence identity: 83
5cyt R - percentage of sequence identity: 62