
|

|

|
| Information on 1vjs |
| PDB: 1vjs Compound: alpha-amylase Classification: HYDROLASE Entry date in PDB: 1997-03-12 Resolution [Å]: 1.70 R-Factor: 0.199 |
CHAIN: - SWISS-PROT/TREMBL:
P06278
KEYWORD: 3D-structure Calcium-binding Carbohydrate metabolism Glycosidase Hydrolase Signal EC: 3.2.1.1 SCOP: b.71.1.1 All beta proteins Glycosyl hydrolase domain Glycosyl hydrolase domain alpha-Amylases, C-terminal beta-sheet domain Bacterial alpha-Amylase Bacillus licheniformis SCOP: c.1.8.1 Alpha and beta proteins (a/b) TIM beta/alpha-barrel (Trans)glycosidases Amylase, catalytic domain Bacterial alpha-amylase Bacillus licheniformis GO: alpha-amylase activity carbohydrate metabolism |
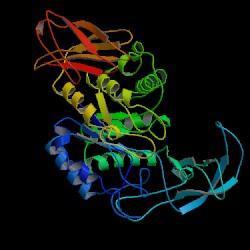
Image Source: PDB |
| Stored Loops of 1vjs |
Loops in ArchDB-EC clusters
1vjs_*_312 - HE => SUBCLASS : 4.10.1
Loops not clustered in ArchDB
1vjs_*_467 - AR
1vjs_*_460 - AR
1vjs_*_434 - AR
1vjs_*_424 - AR
1vjs_*_175 - AR
1vjs_*_133 - AR
1vjs_*_105 - AR
1vjs_*_96 - AR
1vjs_*_7 - EH
1vjs_*_282 - EH
1vjs_*_39 - EH
1vjs_*_320 - EH
1vjs_*_199 - EH
1vjs_*_227 - EH
1vjs_*_360 - EH
1vjs_*_257 - EH
1vjs_*_160 - HA
1vjs_*_399 - HA
1vjs_*_408 - HA
1vjs_*_448 - HA
1vjs_*_111 - HA
1vjs_*_80 - HE
1vjs_*_206 - HE
1vjs_*_238 - HE
1vjs_*_267 - HE
1vjs_*_344 - HE
1vjs_*_381 - HE
1vjs_*_29 - HE
1vjs_*_286 - HH
1vjs_*_363 - HH
1vjs_*_21 - HH
| Homologous structures to 1vjs classified in ArchDB |
1bli - - percentage of sequence identity: 98
1bpl A - percentage of sequence identity: 100
1bpl B - percentage of sequence identity: 100
1e3x A - percentage of sequence identity: 86
1e3z A - percentage of sequence identity: 86
1e40 A - percentage of sequence identity: 86
1ud3 A - percentage of sequence identity: 62
1ud4 A - percentage of sequence identity: 62
1ud5 A - percentage of sequence identity: 62
1ud6 A - percentage of sequence identity: 62
1ud8 A - percentage of sequence identity: 62
1w9x A - percentage of sequence identity: 70
1wp6 A - percentage of sequence identity: 67
1wpc A - percentage of sequence identity: 67