
|

|

|
| Information on 1jbo |
| PDB: 1jbo Compound: c-phycocyanin alpha chain Classification: PHOTOSYNTHESIS Entry date in PDB: 2003-03-18 Resolution [Å]: 1.45 R-Factor: 0.146 |
CHAIN: A SWISS-PROT/TREMBL:
P50032
KEYWORD: 3D-structure Bile pigment Complete proteome Electron transport Photosynthesis Phycobilisome SCOP: a.1.1.3 All alpha proteins Globin-like Globin-like Phycocyanin-like phycobilisome proteins Phycocyanin alpha subunit Synechococcus elongatus GO: light-harvesting complex (sensu Viridiplantae) phycobilisome electron transport photosynthesis CHAIN: B SWISS-PROT/TREMBL:
P50033
KEYWORD: 3D-structure Bile pigment Complete proteome Electron transport Methylation Photosynthesis Phycobilisome SCOP: a.1.1.3 All alpha proteins Globin-like Globin-like Phycocyanin-like phycobilisome proteins Phycocyanin beta subunit Synechococcus elongatus GO: light-harvesting complex (sensu Viridiplantae) phycobilisome electron transport mannitol transport photosynthesis |
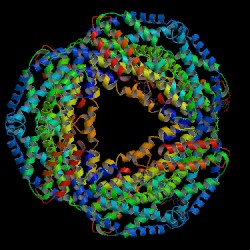
Image Source: PDB |
| Stored Loops of 1jbo |
Loops in ArchDB40 clusters
1jbo_B_78 - HH => SUBCLASS : 2.4.1
1jbo_A_78 - HH => SUBCLASS : 2.4.1
1jbo_B_105 - HH => SUBCLASS : 2.10.1
1jbo_A_105 - HH => SUBCLASS : 2.10.1
1jbo_B_115 - HH => SUBCLASS : 3.2.3
1jbo_A_115 - HH => SUBCLASS : 3.2.3
1jbo_B_115 - HH => SUBCLASS : 3.3.5
1jbo_B_4 - HH => SUBCLASS : 5.3.3
Loops in ArchDB95 clusters
1jbo_B_78 - HH => SUBCLASS : 2.5.1
1jbo_A_78 - HH => SUBCLASS : 2.5.2
1jbo_B_78 - HH => SUBCLASS : 2.5.2
1jbo_B_105 - HH => SUBCLASS : 2.9.1
1jbo_A_105 - HH => SUBCLASS : 2.9.1
1jbo_A_115 - HH => SUBCLASS : 3.1.4
1jbo_A_115 - HH => SUBCLASS : 3.2.5
1jbo_B_115 - HH => SUBCLASS : 3.2.5
1jbo_A_115 - HH => SUBCLASS : 3.4.4
1jbo_B_115 - HH => SUBCLASS : 3.4.9
1jbo_B_4 - HH => SUBCLASS : 5.1.2
1jbo_A_4 - HH => SUBCLASS : 5.7.2
Loops not clustered in ArchDB
1jbo_A_21 - HH
1jbo_A_48 - HH
1jbo_A_126 - HH
1jbo_B_21 - HH
1jbo_B_34 - HH
1jbo_B_48 - HH
1jbo_B_126 - HH
| Homologous structures to 1jbo classified in ArchDB |
1cpc A - percentage of sequence identity: 73
1cpc B - percentage of sequence identity: 80
1cpc K - percentage of sequence identity: 73
1f99 K - percentage of sequence identity: 77
1f99 M - percentage of sequence identity: 77
1gh0 A - percentage of sequence identity: 76
1gh0 B - percentage of sequence identity: 81
1gh0 C - percentage of sequence identity: 76
1gh0 E - percentage of sequence identity: 76
1gh0 G - percentage of sequence identity: 76
1gh0 I - percentage of sequence identity: 76
1gh0 K - percentage of sequence identity: 76
1gh0 M - percentage of sequence identity: 76
1gh0 O - percentage of sequence identity: 76
1gh0 Q - percentage of sequence identity: 76
1gh0 S - percentage of sequence identity: 76
1gh0 U - percentage of sequence identity: 76
1gh0 W - percentage of sequence identity: 76
1ha7 A - percentage of sequence identity: 75
1ha7 C - percentage of sequence identity: 75
1ha7 E - percentage of sequence identity: 75
1ha7 G - percentage of sequence identity: 75
1ha7 I - percentage of sequence identity: 75
1ha7 K - percentage of sequence identity: 75
1ha7 M - percentage of sequence identity: 75
1ha7 O - percentage of sequence identity: 75
1ha7 Q - percentage of sequence identity: 75
1ha7 S - percentage of sequence identity: 75
1ha7 U - percentage of sequence identity: 75
1ha7 W - percentage of sequence identity: 75
1i7y A - percentage of sequence identity: 99
1ktp A - percentage of sequence identity: 99
1on7 A - percentage of sequence identity: 99
2c7j A - percentage of sequence identity: 68
2c7k A - percentage of sequence identity: 68
2c7l A - percentage of sequence identity: 68