| Home | Trees | Indices | Help |
|
|---|
|
|

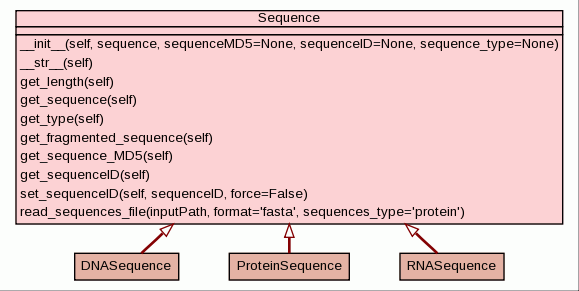
Class to represent a biologic sequencial entity, as nucleotide sequences, protein sequences, etc
This class is suposed to be an abstract class. Only instances for their subclasses should be created
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
|||
|
Inherited from |
|||
|
|||
|
|||
|
|||
|
Inherited from |
|||
|
|||
"sequence": the sequence itself. It is processed to eliminate non sequence elements "sequenceMD5" is the digested md5 code of the sequence. It usually has not to be defined as a parameter, as it implements a method to calculate it "sequenceID" is the unique identifier for the sequence. A sequence must have a sequenceID only when it has been inserted into database "sequence_type" is the type of sequence (protein, dna or rna)
|
str(x)
|
Now: SequenceMD5 is used only for internal working, and it is represented in a 16-byte string Previous: Return MD5 code for sequence "sequence" (MD5 hexdigestion of sequence + its leading 4 chars + its last 4 chars) |
Sets the sequenceID for a sequence. If it contains a sequence ID it should not be modified. In special cases where it should be able to be modified, use the parameter force=True |
Returns a list of sequence objects contained in a file in the specified format "format" can be: fasta "sequences_type" can be: protein, dna or rna |
| Home | Trees | Indices | Help |
|
|---|
| Generated by Epydoc 3.0.1 on Tue May 26 17:06:27 2009 | http://epydoc.sourceforge.net |