
|

|

|
| Information on 1a12 |
| PDB: 1a12 Compound: regulator of chromosome condensation 1 Classification: GUANINE NUCLEOTIDE EXCHANGE FACTOR Entry date in PDB: 1999-01-13 Resolution [Å]: 1.70 R-Factor: 0.189 |
CHAIN: A SWISS-PROT/TREMBL:
P18754
KEYWORD: 3D-structure Cell cycle DNA-binding Guanine-nucleotide releasing factor Mitosis Nuclear protein Repeat SCOP: b.69.5.1 All beta proteins 7-bladed beta-propeller RCC1/BLIP-II Regulator of chromosome condensation RCC1 Regulator of chromosome condensation RCC1 Human (Homo sapiens) GO: CHAIN: B SWISS-PROT/TREMBL:
P18754
KEYWORD: 3D-structure Cell cycle DNA-binding Guanine-nucleotide releasing factor Mitosis Nuclear protein Repeat SCOP: b.69.5.1 All beta proteins 7-bladed beta-propeller RCC1/BLIP-II Regulator of chromosome condensation RCC1 Regulator of chromosome condensation RCC1 Human (Homo sapiens) GO: CHAIN: C SWISS-PROT/TREMBL:
P18754
KEYWORD: 3D-structure Cell cycle DNA-binding Guanine-nucleotide releasing factor Mitosis Nuclear protein Repeat SCOP: b.69.5.1 All beta proteins 7-bladed beta-propeller RCC1/BLIP-II Regulator of chromosome condensation RCC1 Regulator of chromosome condensation RCC1 Human (Homo sapiens) GO: |
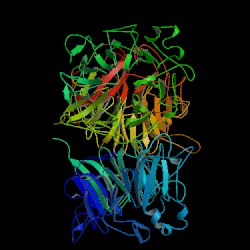
Image Source: PDB |
| Stored Loops of 1a12 |
Loops in ArchDB40 clusters
1a12_A_146 - HA => SUBCLASS : 2.2.2
1a12_A_174 - HA => SUBCLASS : 2.2.3
1a12_A_121 - HA => SUBCLASS : 2.2.3
1a12_A_347 - HA => SUBCLASS : 2.2.3
1a12_A_69 - HA => SUBCLASS : 2.2.3
1a12_A_296 - HA => SUBCLASS : 2.2.3
1a12_A_399 - HA => SUBCLASS : 2.2.3
1a12_A_183 - HA => SUBCLASS : 4.2.1
1a12_A_251 - HA => SUBCLASS : 4.2.1
1a12_A_130 - HA => SUBCLASS : 4.2.1
1a12_A_356 - HA => SUBCLASS : 4.2.1
1a12_A_78 - HA => SUBCLASS : 4.2.1
1a12_A_305 - HA => SUBCLASS : 4.2.1
1a12_A_113 - AR => SUBCLASS : 2.1.2
1a12_A_113 - AR => SUBCLASS : 3.10.1
1a12_A_162 - AR => SUBCLASS : 3.16.2
1a12_A_335 - AR => SUBCLASS : 6.6.2
1a12_A_335 - AR => SUBCLASS : 6.12.1
Loops in ArchDB95 clusters
1a12_A_113 - AR => SUBCLASS : 5.18.1
1a12_A_335 - AR => SUBCLASS : 6.21.1
1a12_A_146 - HA => SUBCLASS : 2.2.2
1a12_A_121 - HA => SUBCLASS : 2.2.3
1a12_A_399 - HA => SUBCLASS : 2.2.3
1a12_A_296 - HA => SUBCLASS : 2.2.3
1a12_A_69 - HA => SUBCLASS : 2.2.3
1a12_A_174 - HA => SUBCLASS : 2.2.3
1a12_A_347 - HA => SUBCLASS : 2.2.3
1a12_A_183 - HA => SUBCLASS : 3.1.2
1a12_A_356 - HA => SUBCLASS : 3.1.2
1a12_A_251 - HA => SUBCLASS : 3.1.2
1a12_A_130 - HA => SUBCLASS : 3.1.2
1a12_A_305 - HA => SUBCLASS : 3.1.2
1a12_A_78 - HA => SUBCLASS : 3.1.2
Loops not clustered in ArchDB
1a12_A_56 - AR
1a12_A_140 - AR
1a12_A_151 - AR
1a12_A_227 - AR
1a12_A_385 - AR
1a12_A_280 - AR
1a12_A_193 - EH
1a12_A_36 - HA
1a12_A_366 - HA
1a12_A_315 - HA
1a12_A_261 - HA
1a12_A_242 - HA
1a12_A_88 - HA
1a12_A_220 - HE
| Homologous structures to 1a12 classified in ArchDB |
1a12 A - percentage of sequence identity: 100
1a12 B - percentage of sequence identity: 100
1a12 C - percentage of sequence identity: 100
1i2m B - percentage of sequence identity: 100
1i2m D - percentage of sequence identity: 100