
|

|

|
| Information on 1d6a |
| PDB: 1d6a Compound: pokeweed antiviral protein Classification: HYDROLASE Entry date in PDB: 1999-12-16 Resolution [Å]: 2.10 R-Factor: 0.190 |
CHAIN: A SWISS-PROT/TREMBL:
P10297
KEYWORD: 3D-structure Antiviral Hydrolase Plant defense Protein synthesis inhibitor Signal Toxin EC: 3.2.2.22 SCOP: d.165.1.1 Alpha and beta proteins (a+b) Ribosome inactivating proteins (RIP) Ribosome inactivating proteins (RIP) Plant cytotoxins Pokeweed antiviral protein alpha Pokeweed (Phytolacca americana) GO: rRNA N-glycosylase activity negative regulation of protein biosynthesis CHAIN: B SWISS-PROT/TREMBL:
P10297
KEYWORD: 3D-structure Antiviral Hydrolase Plant defense Protein synthesis inhibitor Signal Toxin EC: 3.2.2.22 SCOP: d.165.1.1 Alpha and beta proteins (a+b) Ribosome inactivating proteins (RIP) Ribosome inactivating proteins (RIP) Plant cytotoxins Pokeweed antiviral protein alpha Pokeweed (Phytolacca americana) GO: rRNA N-glycosylase activity negative regulation of protein biosynthesis |
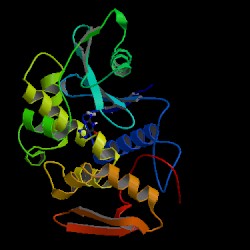
Image Source: PDB |
| Stored Loops of 1d6a |
Loops in ArchDB40 clusters
1d6a_A_33 - HA => SUBCLASS : 2.1.1
1d6a_A_73 - HA => SUBCLASS : 3.4.1
1d6a_A_61 - HA => SUBCLASS : 4.1.2
1d6a_A_222 - HA => SUBCLASS : 4.2.2
1d6a_A_49 - HA => SUBCLASS : 4.5.1
1d6a_A_222 - HA => SUBCLASS : 5.1.5
1d6a_A_3 - EH => SUBCLASS : 6.26.1
1d6a_A_160 - HH => SUBCLASS : 1.2.1
Loops in ArchDB95 clusters
1d6a_A_3 - EH => SUBCLASS : 5.61.1
1d6a_A_112 - EH => SUBCLASS : 7.33.1
1d6a_A_160 - HH => SUBCLASS : 1.2.1
1d6a_A_181 - HH => SUBCLASS : 6.38.1
1d6a_A_49 - HA => SUBCLASS : 3.2.1
1d6a_A_73 - HA => SUBCLASS : 3.5.1
1d6a_A_222 - HA => SUBCLASS : 5.1.1
1d6a_A_222 - HA => SUBCLASS : 5.1.2
1d6a_A_222 - HA => SUBCLASS : 5.1.4
Loops not clustered in ArchDB
1d6a_A_37 - AR
1d6a_A_84 - EH
1d6a_A_236 - EH
1d6a_A_13 - HE
1d6a_A_96 - HE
1d6a_A_208 - HE
1d6a_A_123 - HH
1d6a_A_142 - HH
1d6a_A_199 - HH
| Homologous structures to 1d6a classified in ArchDB |
1apa - - percentage of sequence identity: 74
1d6a A - percentage of sequence identity: 100
1d6a B - percentage of sequence identity: 100
1gik A - percentage of sequence identity: 76
1j1q A - percentage of sequence identity: 76
1j1r A - percentage of sequence identity: 76
1j1s A - percentage of sequence identity: 76
1paf A - percentage of sequence identity: 100
1paf B - percentage of sequence identity: 100
1pag A - percentage of sequence identity: 100
1pag B - percentage of sequence identity: 100
1qcg A - percentage of sequence identity: 100
1qcg B - percentage of sequence identity: 100
1qci A - percentage of sequence identity: 100
1qci B - percentage of sequence identity: 100
1qcj A - percentage of sequence identity: 100
1qcj B - percentage of sequence identity: 100