
|

|

|
| Information on 1erj |
| PDB: 1erj Compound: transcriptional repressor tup1 Classification: TRANSCRIPTION INHIBITOR Entry date in PDB: 2000-07-26 Resolution [Å]: 2.30 R-Factor: 0.228 |
CHAIN: A SWISS-PROT/TREMBL:
P16649
KEYWORD: 3D-structure Repeat Repressor Transcription regulation WD repeat SCOP: b.69.4.1 All beta proteins 7-bladed beta-propeller WD40-repeat WD40-repeat Tup1, C-terminal domain Baker's yeast (Saccharomyces cerevisiae) GO: CHAIN: B SWISS-PROT/TREMBL:
P16649
KEYWORD: 3D-structure Repeat Repressor Transcription regulation WD repeat SCOP: b.69.4.1 All beta proteins 7-bladed beta-propeller WD40-repeat WD40-repeat Tup1, C-terminal domain Baker's yeast (Saccharomyces cerevisiae) GO: CHAIN: C SWISS-PROT/TREMBL:
P16649
KEYWORD: 3D-structure Repeat Repressor Transcription regulation WD repeat SCOP: b.69.4.1 All beta proteins 7-bladed beta-propeller WD40-repeat WD40-repeat Tup1, C-terminal domain Baker's yeast (Saccharomyces cerevisiae) GO: |
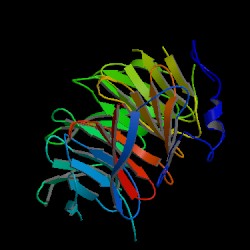
Image Source: PDB |
| Stored Loops of 1erj |
Loops in ArchDB40 clusters
1erj_A_315 - HA => SUBCLASS : 2.2.1
1erj_A_358 - HA => SUBCLASS : 2.6.1
1erj_A_644 - HA => SUBCLASS : 3.1.1
1erj_A_499 - HA => SUBCLASS : 3.1.1
1erj_A_590 - HA => SUBCLASS : 3.1.1
1erj_A_691 - HA => SUBCLASS : 3.1.1
1erj_A_358 - HA => SUBCLASS : 3.7.1
1erj_A_467 - HA => SUBCLASS : 4.1.2
1erj_A_653 - HA => SUBCLASS : 4.1.2
1erj_A_551 - HA => SUBCLASS : 4.1.2
1erj_A_367 - HA => SUBCLASS : 4.1.2
1erj_A_508 - HA => SUBCLASS : 4.1.2
1erj_A_541 - HA => SUBCLASS : 4.2.1
1erj_A_457 - HA => SUBCLASS : 4.2.1
1erj_A_488 - HA => SUBCLASS : 5.2.1
1erj_A_579 - HA => SUBCLASS : 5.2.1
1erj_A_446 - HA => SUBCLASS : 5.2.1
1erj_A_349 - HA => SUBCLASS : 5.2.1
1erj_A_518 - AR => SUBCLASS : 1.1.2
1erj_A_622 - AR => SUBCLASS : 3.2.4
1erj_A_333 - AR => SUBCLASS : 3.2.11
1erj_A_664 - AR => SUBCLASS : 3.2.12
1erj_A_664 - AR => SUBCLASS : 5.1.2
1erj_A_476 - AR => SUBCLASS : 5.23.1
1erj_A_333 - AR => SUBCLASS : 5.25.1
Loops in ArchDB95 clusters
1erj_A_518 - AR => SUBCLASS : 3.1.2
1erj_A_622 - AR => SUBCLASS : 3.1.9
1erj_A_518 - AR => SUBCLASS : 3.1.12
1erj_A_333 - AR => SUBCLASS : 3.1.19
1erj_A_664 - AR => SUBCLASS : 3.4.7
1erj_A_333 - AR => SUBCLASS : 5.3.2
1erj_A_476 - AR => SUBCLASS : 5.25.1
1erj_A_664 - AR => SUBCLASS : 5.54.1
1erj_A_315 - HA => SUBCLASS : 2.2.1
1erj_A_358 - HA => SUBCLASS : 2.4.2
1erj_A_691 - HA => SUBCLASS : 3.1.1
1erj_A_644 - HA => SUBCLASS : 3.1.1
1erj_A_499 - HA => SUBCLASS : 3.1.1
1erj_A_590 - HA => SUBCLASS : 3.1.1
1erj_A_541 - HA => SUBCLASS : 3.1.2
1erj_A_457 - HA => SUBCLASS : 3.1.2
1erj_A_467 - HA => SUBCLASS : 4.1.2
1erj_A_367 - HA => SUBCLASS : 4.1.2
1erj_A_551 - HA => SUBCLASS : 4.1.2
1erj_A_653 - HA => SUBCLASS : 4.1.2
1erj_A_508 - HA => SUBCLASS : 4.1.2
1erj_A_488 - HA => SUBCLASS : 5.2.1
1erj_A_446 - HA => SUBCLASS : 5.2.1
1erj_A_579 - HA => SUBCLASS : 5.2.1
1erj_A_349 - HA => SUBCLASS : 5.2.1
1erj_A_358 - HA => SUBCLASS : 5.3.1
1erj_A_633 - HA => SUBCLASS : 5.23.1
Loops not clustered in ArchDB
1erj_A_322 - AR
1erj_A_377 - AR
1erj_A_561 - AR
1erj_A_529 - HA
1erj_A_599 - HA
1erj_A_675 - HA
1erj_A_300 - HE
| Homologous structures to 1erj classified in ArchDB |
1erj A - percentage of sequence identity: 100
1erj B - percentage of sequence identity: 100
1erj C - percentage of sequence identity: 100