
|

|

|
| Information on 1i4n |
| PDB: 1i4n Compound: indole-3-glycerol phosphate synthase Classification: LYASE Entry date in PDB: 2002-03-20 Resolution [Å]: 2.50 R-Factor: 0.169 |
CHAIN: A SWISS-PROT/TREMBL:
Q56319
KEYWORD: 3D-structure Complete proteome Decarboxylase Lyase Tryptophan biosynthesis EC: 4.1.1.48 SCOP: c.1.2.4 Alpha and beta proteins (a/b) TIM beta/alpha-barrel Ribulose-phoshate binding barrel Tryptophan biosynthesis enzymes Indole-3-glycerophosphate synthase, IPGS Thermotoga maritima GO: indole-3-glycerol-phosphate synthase activity tryptophan metabolism CHAIN: B SWISS-PROT/TREMBL:
Q56319
KEYWORD: 3D-structure Complete proteome Decarboxylase Lyase Tryptophan biosynthesis EC: 4.1.1.48 SCOP: c.1.2.4 Alpha and beta proteins (a/b) TIM beta/alpha-barrel Ribulose-phoshate binding barrel Tryptophan biosynthesis enzymes Indole-3-glycerophosphate synthase, IPGS Thermotoga maritima GO: indole-3-glycerol-phosphate synthase activity tryptophan metabolism |
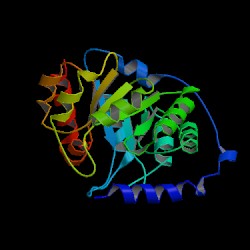
Image Source: PDB |
| Stored Loops of 1i4n |
Loops in ArchDB40 clusters
1i4n_A_153 - EH => SUBCLASS : 2.1.1
1i4n_A_105 - EH => SUBCLASS : 4.15.1
1i4n_A_127 - EH => SUBCLASS : 5.4.1
1i4n_A_77 - EH => SUBCLASS : 9.12.1
1i4n_A_232 - HH => SUBCLASS : 3.6.4
Loops in ArchDB95 clusters
1i4n_A_153 - EH => SUBCLASS : 2.1.2
1i4n_A_105 - EH => SUBCLASS : 4.9.2
1i4n_A_127 - EH => SUBCLASS : 5.2.2
1i4n_A_127 - EH => SUBCLASS : 5.8.4
1i4n_A_77 - EH => SUBCLASS : 9.23.1
1i4n_A_232 - HH => SUBCLASS : 3.5.3
Loops in ArchDB-EC clusters
1i4n_A_232 - HH => SUBCLASS : 3.7.3
Loops not clustered in ArchDB
1i4n_A_43 - EH
1i4n_A_175 - EH
1i4n_A_205 - EH
1i4n_A_227 - EH
1i4n_A_218 - HE
1i4n_A_192 - HE
1i4n_A_161 - HE
1i4n_A_137 - HE
1i4n_A_115 - HE
1i4n_A_92 - HE
1i4n_A_64 - HE
1i4n_A_31 - HE
1i4n_A_4 - HH
| Homologous structures to 1i4n classified in ArchDB |
1i4n A - percentage of sequence identity: 100
1i4n B - percentage of sequence identity: 100
1j5t A - percentage of sequence identity: 99