
|

|

|
| Information on 1mxr |
| PDB: 1mxr Compound: ribonucleotide reductase r2 Classification: OXIDOREDUCTASE Entry date in PDB: 2003-03-25 Resolution [Å]: 1.42 R-Factor: 0.160 |
CHAIN: A SWISS-PROT/TREMBL:
P00453
KEYWORD: 3D-structure Complete proteome DNA replication Iron Metal-binding Oxidoreductase EC: 1.17.4.1 SCOP: a.25.1.2 All alpha proteins Ferritin-like Ferritin-like Ribonucleotide reductase-like Ribonucleotide reductase R2 Escherichia coli GO: ribonucleoside-diphosphate reductase activity deoxyribonucleoside diphosphate metabolism CHAIN: B SWISS-PROT/TREMBL:
P00453
KEYWORD: 3D-structure Complete proteome DNA replication Iron Metal-binding Oxidoreductase EC: 1.17.4.1 SCOP: a.25.1.2 All alpha proteins Ferritin-like Ferritin-like Ribonucleotide reductase-like Ribonucleotide reductase R2 Escherichia coli GO: ribonucleoside-diphosphate reductase activity deoxyribonucleoside diphosphate metabolism |
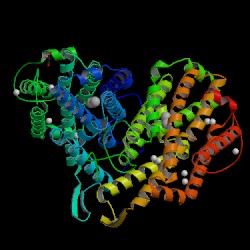
Image Source: PDB |
| Stored Loops of 1mxr |
Loops in ArchDB40 clusters
1mxr_A_180 - EH => SUBCLASS : 1.1.3
1mxr_A_133 - HH => SUBCLASS : 1.1.7
1mxr_A_58 - HH => SUBCLASS : 2.1.1
1mxr_A_58 - HH => SUBCLASS : 2.1.4
1mxr_A_144 - HH => SUBCLASS : 2.13.1
1mxr_A_34 - HH => SUBCLASS : 8.9.1
Loops in ArchDB95 clusters
1mxr_A_180 - EH => SUBCLASS : 1.1.4
1mxr_A_133 - HH => SUBCLASS : 1.1.6
1mxr_A_133 - HH => SUBCLASS : 1.1.10
1mxr_A_58 - HH => SUBCLASS : 2.1.1
1mxr_A_144 - HH => SUBCLASS : 2.9.2
1mxr_A_102 - HH => SUBCLASS : 3.5.7
1mxr_A_207 - HH => SUBCLASS : 3.31.1
1mxr_A_34 - HH => SUBCLASS : 8.9.1
Loops in ArchDB-EC clusters
1mxr_A_58 - HH => SUBCLASS : 2.1.4
Loops not clustered in ArchDB
1mxr_A_173 - HA
1mxr_A_173 - HA
1mxr_A_153 - HE
1mxr_A_67 - HH
1mxr_A_186 - HH
1mxr_A_225 - HH
1mxr_A_259 - HH
1mxr_A_269 - HH
| Homologous structures to 1mxr classified in ArchDB |
1av8 A - percentage of sequence identity: 100
1av8 B - percentage of sequence identity: 100
1biq A - percentage of sequence identity: 99
1biq B - percentage of sequence identity: 99
1jpr A - percentage of sequence identity: 100
1jpr B - percentage of sequence identity: 100
1jqc A - percentage of sequence identity: 100
1jqc B - percentage of sequence identity: 100
1mrr A - percentage of sequence identity: 99
1mrr B - percentage of sequence identity: 99
1mxr A - percentage of sequence identity: 100
1mxr B - percentage of sequence identity: 100
1pfr A - percentage of sequence identity: 99
1pfr B - percentage of sequence identity: 99
1pim A - percentage of sequence identity: 99
1pim B - percentage of sequence identity: 99
1piu A - percentage of sequence identity: 99
1piu B - percentage of sequence identity: 99
1piy A - percentage of sequence identity: 99
1piy B - percentage of sequence identity: 99
1piz A - percentage of sequence identity: 99
1piz B - percentage of sequence identity: 99
1pj0 A - percentage of sequence identity: 99
1pj0 B - percentage of sequence identity: 99
1pj1 A - percentage of sequence identity: 99
1pj1 B - percentage of sequence identity: 99
1pm2 A - percentage of sequence identity: 99
1pm2 B - percentage of sequence identity: 99
1r65 A - percentage of sequence identity: 99
1r65 B - percentage of sequence identity: 99
1rib A - percentage of sequence identity: 99
1rib B - percentage of sequence identity: 99
1rnr A - percentage of sequence identity: 99
1rnr B - percentage of sequence identity: 99
1rsr A - percentage of sequence identity: 99
1rsr B - percentage of sequence identity: 99
1rsv A - percentage of sequence identity: 99
1rsv B - percentage of sequence identity: 99
1xik A - percentage of sequence identity: 100
1xik B - percentage of sequence identity: 100
1yfd A - percentage of sequence identity: 99
1yfd B - percentage of sequence identity: 99
2alx A - percentage of sequence identity: 100
2av8 A - percentage of sequence identity: 99
2av8 B - percentage of sequence identity: 99