
|

|

|
| Information on 1o65 |
| PDB: 1o65 Compound: hypothetical protein yiim Classification: STRUCTURAL GENOMICS, UNKNOWN FUNCTION Entry date in PDB: 2003-11-11 Resolution [Å]: 2.33 R-Factor: 0.231 |
CHAIN: A SWISS-PROT/TREMBL:
P32157
KEYWORD: Complete proteome Hypothetical protein SCOP: b.58.1.2 All beta proteins PK beta-barrel domain-like PK beta-barrel domain-like MOSC (MOCO sulphurase C-terminal) domain Hypothetical protein YiiM Escherichia coli GO: CHAIN: B SWISS-PROT/TREMBL:
P32157
KEYWORD: Complete proteome Hypothetical protein SCOP: b.58.1.2 All beta proteins PK beta-barrel domain-like PK beta-barrel domain-like MOSC (MOCO sulphurase C-terminal) domain Hypothetical protein YiiM Escherichia coli GO: CHAIN: C SWISS-PROT/TREMBL:
P32157
KEYWORD: Complete proteome Hypothetical protein SCOP: b.58.1.2 All beta proteins PK beta-barrel domain-like PK beta-barrel domain-like MOSC (MOCO sulphurase C-terminal) domain Hypothetical protein YiiM Escherichia coli GO: |
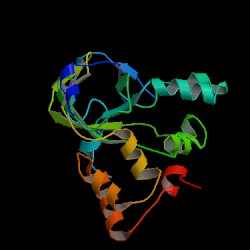
Image Source: PDB |
| Stored Loops of 1o65 |
Loops in ArchDB40 clusters
1o65_A_41 - HA => SUBCLASS : 2.2.2
1o65_A_113 - HA => SUBCLASS : 2.3.1
1o65_A_67 - EH => SUBCLASS : 0.1.6
1o65_A_67 - EH => SUBCLASS : 0.1.53
1o65_A_36 - AR => SUBCLASS : 1.1.8
1o65_A_36 - AR => SUBCLASS : 1.1.10
1o65_A_36 - AR => SUBCLASS : 1.1.17
1o65_A_162 - AR => SUBCLASS : 3.7.1
1o65_A_162 - AR => SUBCLASS : 4.10.3
1o65_A_195 - HH => SUBCLASS : 5.32.1
Loops in ArchDB95 clusters
1o65_A_67 - EH => SUBCLASS : 0.1.5
1o65_A_67 - EH => SUBCLASS : 0.1.49
1o65_A_119 - EH => SUBCLASS : 3.1.6
1o65_A_36 - AR => SUBCLASS : 1.2.4
1o65_A_162 - AR => SUBCLASS : 2.2.1
1o65_A_181 - HH => SUBCLASS : 4.7.2
1o65_A_181 - HH => SUBCLASS : 4.46.1
1o65_A_41 - HA => SUBCLASS : 2.2.2
1o65_A_113 - HA => SUBCLASS : 2.3.1
Loops not clustered in ArchDB
1o65_A_12 - AR
1o65_A_48 - AR
1o65_A_98 - AR
1o65_A_154 - AR
1o65_A_170 - EH
1o65_A_16 - HA
1o65_A_73 - HE
1o65_A_142 - HE
1o65_A_132 - HH
1o65_A_209 - HH
| Homologous structures to 1o65 classified in ArchDB |
1o65 A - percentage of sequence identity: 100
1o65 B - percentage of sequence identity: 100
1o65 C - percentage of sequence identity: 100
1o67 A - percentage of sequence identity: 100
1o67 B - percentage of sequence identity: 100
1o67 C - percentage of sequence identity: 100