
|

|

|
| Information on 1p6o |
| PDB: 1p6o Compound: cytosine deaminase Classification: HYDROLASE Entry date in PDB: 2003-08-19 Resolution [Å]: 1.14 R-Factor: 0.112 |
CHAIN: A SWISS-PROT/TREMBL:
Q12178
KEYWORD: 3D-structure Hydrolase Zinc EC: 3.5.4.1 SCOP: c.97.1.2 Alpha and beta proteins (a/b) Cytidine deaminase-like Cytidine deaminase-like Deoxycytidylate deaminase-like Cytosine deaminase Baker's yeast (Saccharomyces cerevisiae) GO: hydrolase activity zinc ion binding CHAIN: B SWISS-PROT/TREMBL:
Q12178
KEYWORD: 3D-structure Hydrolase Zinc EC: 3.5.4.1 SCOP: c.97.1.2 Alpha and beta proteins (a/b) Cytidine deaminase-like Cytidine deaminase-like Deoxycytidylate deaminase-like Cytosine deaminase Baker's yeast (Saccharomyces cerevisiae) GO: hydrolase activity zinc ion binding |
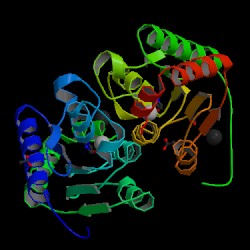
Image Source: PDB |
| Stored Loops of 1p6o |
Loops in ArchDB40 clusters
1p6o_A_118 - HE => SUBCLASS : 1.1.1
1p6o_A_118 - HE => SUBCLASS : 1.1.3
1p6o_A_92 - HE => SUBCLASS : 3.1.1
1p6o_A_11 - HE => SUBCLASS : 4.3.4
1p6o_A_34 - HA => SUBCLASS : 4.1.2
Loops in ArchDB95 clusters
1p6o_A_63 - HH => SUBCLASS : 4.57.1
1p6o_A_118 - HE => SUBCLASS : 1.1.1
1p6o_A_92 - HE => SUBCLASS : 3.1.1
1p6o_A_11 - HE => SUBCLASS : 4.49.1
1p6o_A_34 - HA => SUBCLASS : 4.1.2
Loops in ArchDB-EC clusters
1p6o_A_63 - HH => SUBCLASS : 4.37.1
Loops not clustered in ArchDB
1p6o_A_45 - EH
1p6o_A_82 - EH
1p6o_A_105 - EH
1p6o_A_128 - EH
1p6o_A_34 - HA
1p6o_A_76 - HE
1p6o_A_53 - HH
1p6o_A_135 - HH
| Homologous structures to 1p6o classified in ArchDB |
1ox7 A - percentage of sequence identity: 100
1ox7 B - percentage of sequence identity: 100
1p6o A - percentage of sequence identity: 100
1p6o B - percentage of sequence identity: 100
1rb7 A - percentage of sequence identity: 100
1rb7 B - percentage of sequence identity: 100
1uaq A - percentage of sequence identity: 100
1uaq B - percentage of sequence identity: 100
1ysb A - percentage of sequence identity: 98
1ysb B - percentage of sequence identity: 98
1ysd A - percentage of sequence identity: 98
1ysd B - percentage of sequence identity: 98