
|

|

|
| Information on 1ujk |
| PDB: 1ujk Compound: adp-ribosylation factor binding protein gga1 Classification: PROTEIN TRANSPORT/HYDROLASE Entry date in PDB: 2004-05-11 Resolution [Å]: 1.90 R-Factor: 0.213 |
CHAIN: A SWISS-PROT/TREMBL:
Q9UJY5
KEYWORD: 3D-structure Alternative splicing Coiled coil Golgi stack Protein transport SCOP: a.118.9.2 All alpha proteins alpha-alpha superhelix ENTH/VHS domain VHS domain ADP-ribosylation factor binding protein Gga1 Human (Homo sapiens) GO: clathrin coat of trans-Golgi network vesicle Golgi stack protein transporter activity intra-Golgi transport intracellular protein transport protein complex assembly CHAIN: B SWISS-PROT/TREMBL:
Q9UJY5
KEYWORD: 3D-structure Alternative splicing Coiled coil Golgi stack Protein transport SCOP: a.118.9.2 All alpha proteins alpha-alpha superhelix ENTH/VHS domain VHS domain ADP-ribosylation factor binding protein Gga1 Human (Homo sapiens) GO: clathrin coat of trans-Golgi network vesicle Golgi stack protein transporter activity intra-Golgi transport intracellular protein transport protein complex assembly CHAIN: C SWISS-PROT/TREMBL:
P56817
KEYWORD: 3D-structure Alternative splicing Aspartyl protease Glycoprotein Hydrolase Signal Transmembrane Zymogen EC: 3.4.23.46 GO: CHAIN: D |
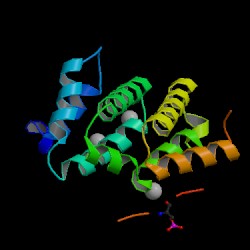
Image Source: PDB |
| Stored Loops of 1ujk |
Loops in ArchDB40 clusters
1ujk_A_79 - HH => SUBCLASS : 1.1.5
1ujk_A_27 - HH => SUBCLASS : 1.2.16
1ujk_A_60 - HH => SUBCLASS : 3.4.1
1ujk_A_43 - HH => SUBCLASS : 3.4.3
1ujk_A_60 - HH => SUBCLASS : 3.4.3
1ujk_A_109 - HH => SUBCLASS : 4.3.1
1ujk_A_10 - HH => SUBCLASS : 8.11.1
Loops in ArchDB95 clusters
1ujk_A_79 - HH => SUBCLASS : 1.1.3
1ujk_A_60 - HH => SUBCLASS : 3.3.1
1ujk_A_43 - HH => SUBCLASS : 3.3.1
1ujk_A_109 - HH => SUBCLASS : 4.2.1
1ujk_A_10 - HH => SUBCLASS : 8.5.1
Loops not clustered in ArchDB
1ujk_A_88 - HH
| Homologous structures to 1ujk classified in ArchDB |
1jpl A - percentage of sequence identity: 72
1jpl B - percentage of sequence identity: 72
1jpl C - percentage of sequence identity: 72
1jpl D - percentage of sequence identity: 72
1juq A - percentage of sequence identity: 72
1juq B - percentage of sequence identity: 72
1juq C - percentage of sequence identity: 72
1juq D - percentage of sequence identity: 72
1jwf A - percentage of sequence identity: 100
1jwg A - percentage of sequence identity: 100
1jwg B - percentage of sequence identity: 100
1lf8 A - percentage of sequence identity: 72
1lf8 B - percentage of sequence identity: 72
1lf8 C - percentage of sequence identity: 72
1lf8 D - percentage of sequence identity: 72
1mhq A - percentage of sequence identity: 63
1mhq B - percentage of sequence identity: 63
1py1 A - percentage of sequence identity: 100
1py1 B - percentage of sequence identity: 100
1py1 C - percentage of sequence identity: 100
1py1 D - percentage of sequence identity: 100
1ujj A - percentage of sequence identity: 100
1ujj B - percentage of sequence identity: 100
1ujk A - percentage of sequence identity: 100
1ujk B - percentage of sequence identity: 100