| Home | Trees | Indices | Help |
|
|---|
|
|

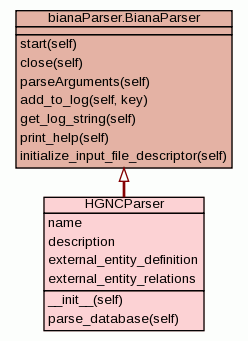
HGNC Parser Class
|
|||
|
|||
|
|||
|
Inherited from Inherited from Inherited from |
|||
|
|||
name = |
|||
description = |
|||
external_entity_definition = |
|||
external_entity_relations = |
|||
|
|||
|
Inherited from |
|||
|
|||
Starts the bianaParser Object
|
Method that implements the specific operations of HGNC parser # Python generated dict 0 : HGNC ID 1 : Approved Symbol 2 : Approved Name 3 : Status 4 : Locus Type 5 : Previous Symbols 6 : Previous Names 7 : Aliases 8 : Name Aliases 9 : Chromosome 10 : Date Approved 11 : Date Modified 12 : Date Symbol Changed 13 : Date Name Changed 14 : Accession Numbers 15 : Enzyme IDs 16 : Entrez Gene ID 17 : Ensembl Gene ID 18 : Mouse Genome Database ID 19 : Specialist Database Links 20 : Specialist Database IDs 21 : Pubmed IDs 22 : RefSeq IDs 23 : Gene Family Name 24 : Record Type 25 : Primary IDs 26 : Secondary IDs 27 : CCDS IDs 28 : VEGA IDs 29 : Locus Specific Databases 30 : GDB ID (mapped data) 31 : Entrez Gene ID (mapped data supplied by NCBI) 32 : OMIM ID (mapped data supplied by NCBI) 33 : RefSeq (mapped data supplied by NCBI) 34 : UniProt ID (mapped data supplied by UniProt) 35 : Ensembl ID (mapped data supplied by Ensembl) 36 : UCSC ID (mapped data supplied by UCSC) 37 : Rat Genome Database ID (mapped data supplied by RGD)
|
|
|||
description
|
external_entity_definition
|
| Home | Trees | Indices | Help |
|
|---|
| Generated by Epydoc 3.0.1 on Tue May 26 17:06:31 2009 | http://epydoc.sourceforge.net |