
|

|

|
| Information on 1pym |
| PDB: 1pym Compound: phosphoenolpyruvate mutase Classification: PHOSPHOTRANSFERASE Entry date in PDB: 1999-07-21 Resolution [Å]: 1.80 R-Factor: 0.188 |
CHAIN: A SWISS-PROT/TREMBL:
P56839
KEYWORD: 3D-structure Isomerase Magnesium EC: 5.4.2.9 SCOP: c.1.12.7 Alpha and beta proteins (a/b) TIM beta/alpha-barrel Phosphoenolpyruvate/pyruvate domain Phosphoenolpyruvate mutase/Isocitrate lyase-like Phosphoenolpyruvate mutase Blue mussel (Mytilus edulis) GO: catalytic activity metabolism CHAIN: B SWISS-PROT/TREMBL:
P56839
KEYWORD: 3D-structure Isomerase Magnesium EC: 5.4.2.9 SCOP: c.1.12.7 Alpha and beta proteins (a/b) TIM beta/alpha-barrel Phosphoenolpyruvate/pyruvate domain Phosphoenolpyruvate mutase/Isocitrate lyase-like Phosphoenolpyruvate mutase Blue mussel (Mytilus edulis) GO: catalytic activity metabolism |
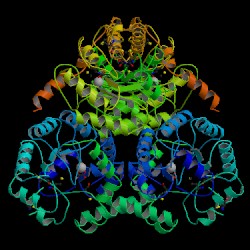
Image Source: PDB |
| Stored Loops of 1pym |
Loops in ArchDB-EC clusters
1pym_A_272 - HH => SUBCLASS : 1.3.1
Loops not clustered in ArchDB
1pym_A_43 - EH
1pym_A_82 - EH
1pym_A_110 - EH
1pym_A_155 - EH
1pym_A_186 - EH
1pym_A_169 - HE
1pym_A_135 - HE
1pym_A_93 - HE
1pym_A_64 - HE
1pym_A_29 - HE
1pym_A_7 - HH
1pym_A_47 - HH
1pym_A_197 - HH
1pym_A_224 - HH
1pym_A_240 - HH
1pym_A_263 - HH
| Homologous structures to 1pym classified in ArchDB |
1m1b A - percentage of sequence identity: 100
1m1b B - percentage of sequence identity: 100
1pym A - percentage of sequence identity: 100
1pym B - percentage of sequence identity: 100
1s2t A - percentage of sequence identity: 100
1s2t B - percentage of sequence identity: 100
1s2u A - percentage of sequence identity: 99
1s2u B - percentage of sequence identity: 99
1s2v A - percentage of sequence identity: 100
1s2v B - percentage of sequence identity: 100
1s2v C - percentage of sequence identity: 100
1s2v D - percentage of sequence identity: 100
1s2w A - percentage of sequence identity: 100