
|

|

|
| Information on 1upm |
| PDB: 1upm Compound: ribulose bisphosphate carboxylase large chain Classification: LYASE Entry date in PDB: 2003-10-14 Resolution [Å]: 2.30 R-Factor: 0.230 |
CHAIN: B SWISS-PROT/TREMBL:
P00875
KEYWORD: 3D-structure Acetylation Carbon dioxide fixation Chloroplast Lyase Magnesium Metal-binding Monooxygenase Oxidoreductase Photorespiration Photosynthesis EC: 4.1.1.39 GO: ribulose bisphosphate carboxylase complex (sensu Magnoliophyta) ribulose-bisphosphate carboxylase activity carbon utilization by fixation of carbon dioxide CHAIN: C SWISS-PROT/TREMBL:
Q43832
KEYWORD: 3D-structure Carbon dioxide fixation Chloroplast Lyase Monooxygenase Multigene family Oxidoreductase Photorespiration Photosynthesis Transit peptide EC: 4.1.1.39 GO: ribulose bisphosphate carboxylase complex (sensu Magnoliophyta) ribulose-bisphosphate carboxylase activity carbon utilization by fixation of carbon dioxide CHAIN: E SWISS-PROT/TREMBL:
P00875
KEYWORD: 3D-structure Acetylation Carbon dioxide fixation Chloroplast Lyase Magnesium Metal-binding Monooxygenase Oxidoreductase Photorespiration Photosynthesis EC: 4.1.1.39 GO: ribulose bisphosphate carboxylase complex (sensu Magnoliophyta) ribulose-bisphosphate carboxylase activity carbon utilization by fixation of carbon dioxide CHAIN: F SWISS-PROT/TREMBL:
Q43832
KEYWORD: 3D-structure Carbon dioxide fixation Chloroplast Lyase Monooxygenase Multigene family Oxidoreductase Photorespiration Photosynthesis Transit peptide EC: 4.1.1.39 GO: ribulose bisphosphate carboxylase complex (sensu Magnoliophyta) ribulose-bisphosphate carboxylase activity carbon utilization by fixation of carbon dioxide CHAIN: H SWISS-PROT/TREMBL:
P00875
KEYWORD: 3D-structure Acetylation Carbon dioxide fixation Chloroplast Lyase Magnesium Metal-binding Monooxygenase Oxidoreductase Photorespiration Photosynthesis EC: 4.1.1.39 GO: ribulose bisphosphate carboxylase complex (sensu Magnoliophyta) ribulose-bisphosphate carboxylase activity carbon utilization by fixation of carbon dioxide CHAIN: I SWISS-PROT/TREMBL:
Q43832
KEYWORD: 3D-structure Carbon dioxide fixation Chloroplast Lyase Monooxygenase Multigene family Oxidoreductase Photorespiration Photosynthesis Transit peptide EC: 4.1.1.39 GO: ribulose bisphosphate carboxylase complex (sensu Magnoliophyta) ribulose-bisphosphate carboxylase activity carbon utilization by fixation of carbon dioxide CHAIN: K SWISS-PROT/TREMBL:
P00875
KEYWORD: 3D-structure Acetylation Carbon dioxide fixation Chloroplast Lyase Magnesium Metal-binding Monooxygenase Oxidoreductase Photorespiration Photosynthesis EC: 4.1.1.39 GO: ribulose bisphosphate carboxylase complex (sensu Magnoliophyta) ribulose-bisphosphate carboxylase activity carbon utilization by fixation of carbon dioxide CHAIN: L SWISS-PROT/TREMBL:
P00875
KEYWORD: 3D-structure Acetylation Carbon dioxide fixation Chloroplast Lyase Magnesium Metal-binding Monooxygenase Oxidoreductase Photorespiration Photosynthesis EC: 4.1.1.39 GO: ribulose bisphosphate carboxylase complex (sensu Magnoliophyta) ribulose-bisphosphate carboxylase activity carbon utilization by fixation of carbon dioxide CHAIN: M SWISS-PROT/TREMBL:
Q43832
KEYWORD: 3D-structure Carbon dioxide fixation Chloroplast Lyase Monooxygenase Multigene family Oxidoreductase Photorespiration Photosynthesis Transit peptide EC: 4.1.1.39 GO: ribulose bisphosphate carboxylase complex (sensu Magnoliophyta) ribulose-bisphosphate carboxylase activity carbon utilization by fixation of carbon dioxide CHAIN: O SWISS-PROT/TREMBL:
P00875
KEYWORD: 3D-structure Acetylation Carbon dioxide fixation Chloroplast Lyase Magnesium Metal-binding Monooxygenase Oxidoreductase Photorespiration Photosynthesis EC: 4.1.1.39 GO: ribulose bisphosphate carboxylase complex (sensu Magnoliophyta) ribulose-bisphosphate carboxylase activity carbon utilization by fixation of carbon dioxide CHAIN: P SWISS-PROT/TREMBL:
Q43832
KEYWORD: 3D-structure Carbon dioxide fixation Chloroplast Lyase Monooxygenase Multigene family Oxidoreductase Photorespiration Photosynthesis Transit peptide EC: 4.1.1.39 GO: ribulose bisphosphate carboxylase complex (sensu Magnoliophyta) ribulose-bisphosphate carboxylase activity carbon utilization by fixation of carbon dioxide CHAIN: R SWISS-PROT/TREMBL:
P00875
KEYWORD: 3D-structure Acetylation Carbon dioxide fixation Chloroplast Lyase Magnesium Metal-binding Monooxygenase Oxidoreductase Photorespiration Photosynthesis EC: 4.1.1.39 GO: ribulose bisphosphate carboxylase complex (sensu Magnoliophyta) ribulose-bisphosphate carboxylase activity carbon utilization by fixation of carbon dioxide CHAIN: S SWISS-PROT/TREMBL:
Q43832
KEYWORD: 3D-structure Carbon dioxide fixation Chloroplast Lyase Monooxygenase Multigene family Oxidoreductase Photorespiration Photosynthesis Transit peptide EC: 4.1.1.39 GO: ribulose bisphosphate carboxylase complex (sensu Magnoliophyta) ribulose-bisphosphate carboxylase activity carbon utilization by fixation of carbon dioxide CHAIN: T SWISS-PROT/TREMBL:
Q43832
KEYWORD: 3D-structure Carbon dioxide fixation Chloroplast Lyase Monooxygenase Multigene family Oxidoreductase Photorespiration Photosynthesis Transit peptide EC: 4.1.1.39 GO: ribulose bisphosphate carboxylase complex (sensu Magnoliophyta) ribulose-bisphosphate carboxylase activity carbon utilization by fixation of carbon dioxide CHAIN: V SWISS-PROT/TREMBL:
P00875
KEYWORD: 3D-structure Acetylation Carbon dioxide fixation Chloroplast Lyase Magnesium Metal-binding Monooxygenase Oxidoreductase Photorespiration Photosynthesis EC: 4.1.1.39 GO: ribulose bisphosphate carboxylase complex (sensu Magnoliophyta) ribulose-bisphosphate carboxylase activity carbon utilization by fixation of carbon dioxide CHAIN: W SWISS-PROT/TREMBL:
Q43832
KEYWORD: 3D-structure Carbon dioxide fixation Chloroplast Lyase Monooxygenase Multigene family Oxidoreductase Photorespiration Photosynthesis Transit peptide EC: 4.1.1.39 GO: ribulose bisphosphate carboxylase complex (sensu Magnoliophyta) ribulose-bisphosphate carboxylase activity carbon utilization by fixation of carbon dioxide |
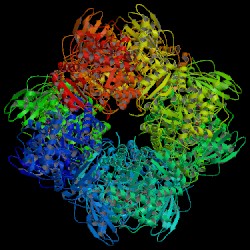
Image Source: PDB |
| Homologous structures to 1upm classified in ArchDB |
1ir1 S - percentage of sequence identity: 94
1rbl A - percentage of sequence identity: 82
3rub S - percentage of sequence identity: 76
8ruc A - percentage of sequence identity: 100