
|

|

|
| Information on 4htc |
| PDB: 4htc Compound: Alpha-thrombin complex with recombinant hirudin (v Classification: HYDROLASE(SERINE PROTEASE) Entry date in PDB: 1994-01-31 Resolution [Å]: 2.30 R-Factor: 0.173 |
CHAIN: H SWISS-PROT/TREMBL:
P00734
KEYWORD: 3D-structure Acute phase Blood coagulation Calcium-binding Disease mutation Gamma-carboxyglutamic acid Glycoprotein Hydrolase Kringle Plasma Polymorphism Repeat Serine protease Signal Vitamin K Zymogen EC: 3.4.21.5 SCOP: b.47.1.2 All beta proteins Trypsin-like serine proteases Trypsin-like serine proteases Eukaryotic proteases Thrombin Human (Homo sapiens) GO: extracellular region calcium ion binding chymotrypsin activity thrombin activity trypsin activity blood coagulation proteolysis and peptidolysis CHAIN: I SWISS-PROT/TREMBL:
P09945
KEYWORD: 3D-structure Multigene family Serine protease inhibitor Signal Sulfation SCOP: g.3.15.2 Small proteins Knottins (small inhibitors, toxins, lectins) Leech antihemostatic proteins Hirudin-like Hirudin Leech (Hirudo medicinalis) GO: serine-type endopeptidase inhibitor activity CHAIN: L SWISS-PROT/TREMBL:
P00734
KEYWORD: 3D-structure Acute phase Blood coagulation Calcium-binding Disease mutation Gamma-carboxyglutamic acid Glycoprotein Hydrolase Kringle Plasma Polymorphism Repeat Serine protease Signal Vitamin K Zymogen EC: 3.4.21.5 SCOP: b.47.1.2 All beta proteins Trypsin-like serine proteases Trypsin-like serine proteases Eukaryotic proteases Thrombin Human (Homo sapiens) GO: extracellular region calcium ion binding chymotrypsin activity thrombin activity trypsin activity blood coagulation proteolysis and peptidolysis |
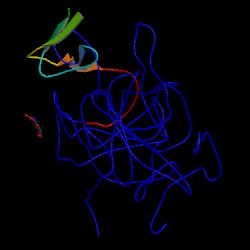
Image Source: PDB |
| Stored Loops of 4htc |
Loops in ArchDB40 clusters
4htc_I_11 - AR => SUBCLASS : 1.2.2
Loops in ArchDB95 clusters
4htc_I_11 - AR => SUBCLASS : 1.1.5
Loops not clustered in ArchDB
4htc_I_21 - AR
4htc_I_38 - AR
4htc_I_14 - HA
4htc_I_26 - HA
| Homologous structures to 4htc classified in ArchDB |
1etr H - percentage of sequence identity: 87
1h8d H - percentage of sequence identity: 97
1hic - - percentage of sequence identity: 86
1hrt I - percentage of sequence identity: 87
1jou B - percentage of sequence identity: 99
1ucy H - percentage of sequence identity: 85
1vr1 H - percentage of sequence identity: 96
2hir - - percentage of sequence identity: 87
2hnt F - percentage of sequence identity: 100
3htc I - percentage of sequence identity: 100
4hir - - percentage of sequence identity: 86
5hir - - percentage of sequence identity: 87
6hir - - percentage of sequence identity: 86
null nu - percentage of sequence identity: 0